Learning objectives
Magnetic Resonance Spectroscopic Imaging (MRSI) reveals biomarkers (metabolites) within biological tissue in a non-invasive and radiation-free way.
This may yield decisive information for posing a diagnosis,
without the need for invasive biopsies.
There are some impediments for MRSI to enter the clinical practice,
among which we focus here on its low sensitivity,
which translates into a low spatial resolution of MRSI compared to MRI.
This drawback is one of several challenges addressed by the new European project TRANSACT,
which aims at transforming MR spectroscopy into...
Background
MRSI combines the principles of MR imaging and MR spectroscopy for visualizing metabolic presence in 2 or 3 spatial dimensions.
Metabolite concentrations are about 10000 times smaller than the water concentration,
which leads to an intrinsically low sensitivity of MRSI.
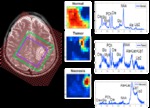
This is usually compensated by lowering the spatial resolution (see Fig.
1),
so that the measurement time can still be acceptable for patients.
By automatic classification,
trained on a large number of example spectra from various brain tumors,
the image can be segmented into different...
Imaging findings OR Procedure details
Spatial resolution of maps derived from MRSI data can be increased artificially by mathematical interpolation.
As a first possibility,
standard 2D interpolation can be performed on each of the metabolite maps of interest.
For instance,
the resolution of the metabolic maps in Figure 1 could be artificially increased such that it matches the MRI resolution.
This can be performed by traditional mathematical tools such as bilinear interpolation or bicubic spline interpolation.
Although this could lead to a visual enhancement of the metabolite maps,
it adds...
Conclusion
Complementing MRSI with information from conventional or advanced MR imaging,
and,
in particular,
increasing spatial resolution of metabolic images,
is bound to have a positive impact on the use of MRSI in clinical practice.
References
[1] Croitor Sava A.R.,
Sima D.M.,
Poullet J.B.,
Wright A.J.,
Heerschap A.,
Van Huffel S.
Exploiting spatial information to estimate metabolite levels in two-dimensional MRSI of heterogeneous brain lesions.NMR in Biomedicine2011,
24(7):824-835.
[2] Luts J.
Classification of brain tumors based on magnetic resonance spectroscopy,
PhD thesis,
Faculty of Engineering,
KU Leuven (Leuven,
Belgium),
2010.
Personal Information
Diana M.
Sima,
Department of Electrical Engineering,
ESAT-SCD,
KU Leuven,
Belgium,
[email protected]
Dirk Smeets,
Dirk Loeckx,
icoMetrix,
Belgium,
[email protected],
[email protected]

![Fig. 1: Example of an MRSI dataset acquired from a brain tumor patient (meningioma). Top left: example of a short-echo time MR spectrum from normal brain tissue. Top right: Anatomic MR image, overlaid with the MRSI grid. The white rectangle indicates the selected volume of interest. Bottom: metabolic maps corresponding to the volume of interest, quantified from the MR spectra for several metabolites, where red represents high concentration and blue represents low concentration. The scale is metabolite-dependent. (Data from [1].) The low sensitivity and low spatial resolution translate in coarse metabolite maps. Although the presence/absence of a metabolite can be indicative for the tumor type and grade, a higher resolution would be beneficial in order to assess the metabolic content more sensitively, e.g., in the peritumoral region. References: Department of Electrical Engineering, KULeuven, Belgium](https://epos.myesr.org/posterimage/esr/ecr2013/118033/media/484047?maxheight=150&maxwidth=150)
![Fig. 3: Example of nosologic images obtained from MRI segmentation, MRSI interpolation and MRSI classification. Top: anatomic MR images of different contrasts. Bottom: nosologic images with the following color code:
• light blue = white matter
• dark blue = gray matter
• green = CSF
• yellow = grade II glioma
• orange = grade III glioma
• dark red = glioblastoma
• purple = meningioma.
The tumor types were confirmed in these cases by histopathology. (Figure adapted from [2].) References: Department of Electrical Engineering, KULeuven, Belgium](https://epos.myesr.org/posterimage/esr/ecr2013/118033/media/484048?maxheight=150&maxwidth=150)