Aims and objectives
The use of radiomics metrics to support radiological reading is rapidly increasing.
These metrics range from the simple (e.g.
maximum intensity) to the very abstract (e.g.
GCLM_Entropy) with a majority of the more interesting results coming from the more complex ones.
In order to bring meaning to these metrics,
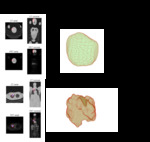
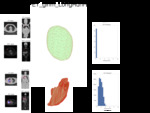
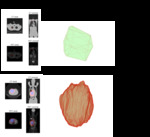
we adopt a visual and comparative approach for examining and interpreting these metrics by applying them to a large body of lung cancer lesions in PET and CT images.
Methods and materials
For this study,
we take a group of 179 patients diagnosed with Lung Cancer with available imaging data.
For each of these patients we take the first PET and CT image.
The lesions within the images are then manually marked using a 3D Slicer-based Annotation Tool and include annotations of Tumor,
Lymph Nodes,
and Metastases following the TNM-atlas.
We take the established library PyRadiomics (radiomics.io) and complement it with a number of other common image features to generate 194 features per lesion covering both PET...
Results
The results for the study are a web-based interface for this database of over 400000 radiomic features covering a range of lesion types from large T4 tumors to inflamed lymph nodes and bone metastases.
These different images and values provide context for understanding the meaning of a given radiomic parameter.
Conclusion
Radiology greatly benefits from the introduction of more reproducible,
quantitative metrics to make reports more detailed and clear for other physicians,
patients,
and clinical trials.
As more and more studies find valuable metrics for predicting disease and outcome,
it will remain important to closely examine what these metrics actually mean for the morphological understanding of the tumor and disease.
Our approach adds understanding to previously large unaccessible quantitative values derived from radiomics approaches and thus paves the way for a broad clinical acceptance of radiomics.
References
Aerts,
H.
J.
W.
L.
et al. (2014) ‘Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach.’,
Nature communications.
Nature Publishing Group,
5,
p.
4006.
doi: 10.1038/ncomms5006.
van Griethuysen,
J.
J.
M.
et al. (2017) ‘Computational Radiomics System to Decode the Radiographic Phenotype.’,
Cancer research.
American Association for Cancer Research,
77(21),
pp.
e104–e107.
doi: 10.1158/0008-5472.CAN-17-0339.
Levy,
M.
A.
et al. (2012) ‘Informatics methods to enable sharing of quantitative imaging research data.’,
Magnetic resonance imaging,
30(9),
pp.
1249–56.
doi: 10.1016/j.mri.2012.04.007.