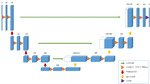
Aims and objectives
Living Donor Liver Transplantation (LDLT) is the commonest form of liver transplantation in Asia.
While Deceasead Donor Liver Transplantation (DDLT) constitutes more than 90% of liver transplantation in the western world,
in India and many other Asian countries,
the majority of transplants performed are LDLT.
Living Donor Liver Transplants is now a well established procedure and has reduced liver transplant waiting list mortality.
The complexities of the procedure,
along with the donor risks,
are the biggest obstacles to widespread use of this valuable treatment option....
Methods and materials
Data Preparation
A test dataset of 15 multi-phasic contrast-enhanced CT scans was provided by Centre for Advanced Research in Imaging,
Neurosciences and Genomics (CARING),
New Delhi,
India.
Exclusion Criteriaentailed:
Morphologic features of cirrhosis,
History of prior liver/biliary surgery or liver tumor ablation procedures
One or more liver lesions greater than 3 cm in size identified by CT or MRI
Portal or hepatic vein thrombosis.
All studies were acquired on a 128-MDCT GE Discovery IQ scanner.
The images were acquired using a matrix size of 512...
Results
Consistency in Liver Volumes
In Fig.
4,
we can see the volumes (in ml) obtained for the two setups.
The liver volumes for the studies are consistent over the two setups with a maximum variation of 2.3% and an average variation of 0.9%.
Duration of Segmentation : Manual vs Automated
In Fig.
3,
we can see the time taken (in mins) for performing liver volumetry on the two setups.
Automation of liver volumetry accelerated the post processing significantly.
Automated Liver volumetry on PredibleLiver takes an...
Conclusion
The study shows that liver volumetry post processing can be significantly accelerated by initializing with Deep Learning based segmentation.
We compared two setups : (1) Commercial CT Volume Viewer and (2) PredibleLiver.
We foundPredibleLiver to require lesser time inperforming volumetric assessment over15 studies as the segmentations comepre-initalized using Deep Learning.
Personal information
Please do not hesitate to reach out to us in case you have any questions or comments:
Vasantha Venugopal,
MD
Centre for Advanced Research in Imaging,
Neuroscience & Genomics,
(CARING),
Mahajan Imaging,
E-19,
Defence Colony,
New Delhi,
INDIA
91 9871438999
[email protected]
www.caring-research.com
Abhijit Chunduru,
BTech
Predible Health
IKP Eden,
#16,
Bhuvanappa Layout,
Adugodi,
Bangalore,
INDIA
+91 9790742906
[email protected]
www.prediblehealth.com
References
1.
Szegedy,
C.,
Vanhoucke,
V.,
Ioffe,
S.,
Shlens,
J.,
Wojna,
Z.: Rethinking the inception architecture for computer vision.
CoRR abs/1512.00567 (2015)
2.
Ren,
S.,
He,
K.,
Girshick,
R.,
Sun,
J.: Faster R-CNN: Towards real-time object detection with region proposal networks.
In: NIPS.
(2015)
3.
Long,
J.,
Shelhamer,
E.,
Darrell,
T.: Fully convolutional networks for semantic segmentation.
In: Proc.
CVPR.
pp.
3431–3440 (2015)
4.
Chen,
L.C.,
Papandreou,
G.,
Kokkinos,
I.,
Murphy,
K.,
Yuille,
A.L.: Semantic image segmentation with deep convolutional nets and fully connected crfs.
In:...