Keywords:
Cardiac, Computer applications, Haematologic, MR, Segmentation, Image registration, Image verification, Haematologic diseases
Authors:
P. Triadyaksa, M. Oudkerk, P. E. Sijens; Groningen/NL
Methods and Materials
Two observers drew free-hand left ventricle epicard and endocard contours on a single image (method 1) and on a CNR-optimized composite image (method 2) derived from multi gradient echo T2* scans acquired with 8 TEs (2.59 - 18.20 ms,
2.23 ms increment) on 36 slices (7 apical,
29 mid or basal) from 21 patients (9 hematological and 12 suspected of cardiomyopathy).
Time delay of drawing between the two methods was more than one week.
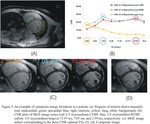
Method 2 images were combinations of those three original images with maximum CNR between myocardium and its surroundings (left and right ventricular blood pool,
lung) as shown in figure 2.
The two methods were assessed by using the same custom-written contour drawing software with the same window-level and window-width setting.
Contour similarities between observers in the two methods were assessed by dice similarity coefficient (DSC) where a minimum value of 0 shows no spatial overlap between two contours and a maximum value of 1 shows complete overlap [8].
DSC results of the two methods are presented as medians ± median absolute deviations and compared by using paired Wilcoxon test.
The inter-observer reproducibility of segmental AHA T2* [1,
9] was assessed on the septum (7 apical septal + 29 x 2 [anteroseptal and inferoseptal] =65) by Bland-Altman analysis [10] with subgroups comparison of minimum segmental T2* value of 20 ms [11].
Bland-Altman results were presented as means ± standard deviations with agreement was presented as limit of agreement.
Correlation of T2* between observers in each method was assessed by Pearson correlation coefficient.
Statistical analyses were performed using IBM SPSS Statistics software version 20 (IBM Corporation,
Somers,
NY,
USA) with P < 0.05 was considered statistically significant.