This poster is published under an
open license. Please read the
disclaimer for further details.
Keywords:
Oncology, Computer applications, eHealth, CT, PACS, RIS, Structured reporting, Computer Applications-Detection, diagnosis, Outcomes analysis, Cancer, Metastases, Pathology
Authors:
D. J. Vining1, C. Popovici2, A. Pitici2, U. I. S. E. N. Salem1, A. Prisacariu2, M. Jurca2, R. Rosu2, A. Tsimberidou1; 1Houston, TX/US, 2Chapel Hill, NC/US
DOI:
10.1594/ecr2014/C-2132
Methods and materials
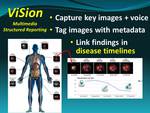
We developed the ViSion multimedia structured reporting system which tags key images with metadata to describe diagnostic findings (3).
ViSion comprises a client-server software arrangement.
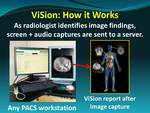
The client resides on a computer workstation and runs in parallel with any vendor’s picture archiving and communication system (PACS) or advance 3D imaging system.
As a radiologist analyzes images with the PACS or 3D imaging system,
the radiologist can press a button on a keyboard or speech microphone to initiate a screen capture of the image finding that is transmitted to the ViSion server hosted on the cloud (Figure 1).
As the radiologist dictates a verbal description of the finding,
that description is also transmitted to the cloud and associated with the key image (Figure 2).
Metadata describing the anatomical location,
radiological observation or diagnosis,
and other salient secondary features of the finding can be extracted from the verbal description using natural language processing techniques or manually using pull-down menus (Figure 3).
The metadata used to label image findings with structured information is managed by an integrated ViSion ontology (i.e.,
controlled vocabulary with defined relationships between terms),
and this ontology currently contains 921 anatomical locations and 12,056 observations/diagnoses.
This ontology enables data mining functions.
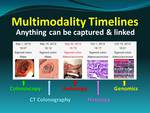
Findings from serial imaging examinations,
including pathology and genomic analysis,
can be linked in graphical disease timelines to illustrate disease progression in terms of images and graphed metrics (Figures 4,
5).
In addition,
we developed data mining tools that can search for disease patterns in the multidisciplinary disease timelines to automate the American Joint Committee on Cancer’s tumor-metastasis-node (TMN) staging of individual patients,
as well as aggregate data from multiple patients enrolled in clinical trials to generate waterfall plots and related statistics (Figures 6,
7).