Type:
Educational Exhibit
Keywords:
Tissue characterisation, Neoplasia, Cancer, Computer Applications-Detection, diagnosis, Computer Applications-3D, CT-Quantitative, CT, Thorax, Lung
Authors:
A. Ottavianelli1, G. Cicchetti1, A. Farchione1, A. R. Larici1, G. Sica1, L. Calandriello2, V. Valentini1, L. Bonomo1, R. Manfredi1; 1Rome/IT, 2Roma/IT
DOI:
10.1594/ecr2018/C-2329
Background
Tumour’s stage is the most important prognosticator used in the management of lung cancer,
anyway even in patient with the same stage a different clinical behaviour has been seen.
Indeed in addition to the anatomic extension of the lesion other factors related to the patient,
tumour histology or “habitat” determine prognosis.
Therefore multifactorial predictive models are mandatory to obtain the so-called “personalized medicine”.
Cancer tissue is represented by a continuous interaction between microenvironment and neoplastic cells,
which are continually forced in genetic mutation to generate an adaptive response to local environment changes.
This process leads to an important genetic and phenotypic heterogeneity of neoplastic tissue [1].
It has been observed that more heterogeneous tumours tend to be more aggressive and to be associated with poorer outcomes,
the last fact often related to poor response to therapy [2].
Investigator have hypothesized that medical image structure is partly determined by tumour histopathologic phenotype and genotype [2-3].
Therefore imaging features can be related to underlie phenotypic and genotypic factors,
as well as the measurements of heterogeneity within medical images could provide further insights into tumour architecture [2-3].
Cancer heterogeneity manifests not only in the spatial dimension (distinct molecular characteristics coexist in the same tumour),
but also in the temporal one (molecular characteristics change over time) [3].
In the clinical practice histology analysis and gene-expression profiling relies on invasive surgical tissue sampling,
whose main drawbacks are that it is not possible to biopsy every part of each tumour at multiple time points [3].
In contrast,
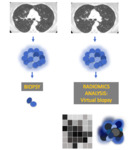
imaging captures intra-tumour heterogeneity in a non-invasive three-dimensional manner,
and can be obtained multiple times as part of routine clinical care [3] (Figure 1).
Radiomics refers to the high-throughput mining of quantitative image features from standard-of-care medical imaging,
their analysis and modelling in relation to prediction targets,
such as clinical end points and genomic features (the so-called Radiogenomics).
Imaging data can be analysed alone or in combination with others “omics” (e.g.
genomics,
metabolomics,
proteomics) or clinical information.
The final aim is to obtain multidisciplinary clinical-decision support systems (CDSS) to improve medical decision-making [3].