Keywords:
MR physics, MR, Imaging sequences, Tissue characterisation
Authors:
D. Cicolari1, D. Lizio2, P. Pedrotti2, R. Sironi2, M. T. Moioli2, A. Lascialfari2, M. Mariani1, A. Torresin2; 1Pavia/IT, 2Milan/IT
DOI:
10.26044/ecr2019/C-1108
Methods and materials
Six vials (ID 2,
4,
11,
13,
14 and 15) of an Eurospin phantom (Diagnostic Sonar,
Livingston,
UK),
filled with agarose gels doped with gadolinium,
were used to compare relaxation times results obtained from measurements performed with different scanners.
Standard reference values of ID inserts were needed to be established with NMR methodology.
First,
the vials were scanned with Tecmag Apollo NMR spectrometer (Tecmag,
Houston,
TX,
USA) at University Physics Department’s laboratories using standard sequences in order to restate reference T1 and T2 relaxation times values at 1.5 T.
Saturation Recovery (SR) and Inversion Recovery (IR) for T1 measurements and Spin Echo (SE) and CPMG (Carr-Purcell-Meiboom-Gill) for T2 measurements were applied.
Sequences were planned with the Tecmag NTNMR software and signals were analyzed with the non-commercial QTNMR software,
developed by NMR-NQR group.
SR and IR data were then fitted with a 3-parameters exponential recovery function [y = A – B exp (- x / T1)]; SE and CPMG data were fitted with a 2-parameters exponential decay function [y = A exp (- x / T2)] [4].
Fitting procedure was performed by means of gnuplot software (http://gnuplot.info).
The vials of the Eurospin phantom were also scanned through two MRI scanners from different vendors: a Siemens MAGNETOM Aera (1.5 T) and a General Electric Signa (1.5 T),
both located in our Hospital.
Phantom images were acquired with body coils and by means of standard clinical sequences SE (Spin Echo,
3 images: TE from 20 ms to 100 ms; TR = 1.5 s) and IR (Inversion Recovery,
8 images: TI from 100 ms to 3300 ms; TR = 5 s).
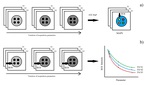
Images were processed with different vendor independent software (Fig.
1): cvi42 (Circle Cardiovascular Imaging Inc.,
http://www.circlecvi.com,
CE mark) and Segment (open source,
[5,
6]); a manual computation of relaxation times was also performed using the gnuplot software.
For each slice,
Cvi42 and Segment generated a map performing a pixel per pixel fit; measurements were executed with ImageJ and Segment by calculating the mean value (and its standard deviation) in a selected ROI on each map.
Then,
a weighted average was taken from relaxation times values measured in corresponding ROIs in maps of each slice.
The intensities,
measured with ImageJ,
in corresponding ROIs selected in each image of the acquisition series were also plotted and fitted by means of gnuplot.
Then,
a weighted average was taken from relaxation times values obtained by the fitting procedure of each slice.