Identifying a Clinical Question and the Research Team
Selection of the patient population and clinical question should be explored between clinical teams and radiologists to ensure appropriate and clinically focused research studies are undertaken.
The research should address an unmet clinical need that aims to bring direct patient benefit.
A multidisciplinary team approach to research utilising skills of radiologists,
clinicians,
physicists,
computer scientists,
data scientists and statisticians is essential and should be identified prior to commencing the study.
Radiologists have the unique position of having an understanding of imaging,
informatics and a clinical background and are ideal research team leaders for radiomic and radiogenomic studies.
Imaging Data
Standardisation of imaging: For optimum results,
radiological studies should be standardised as much as possible.
They should ideally be acquired using the same protocol (resolution,
field of view,
slice thickness,
cut angles,
contrast washout time,
etc.).
Images should be acquired in standardised formats.
Volumetric imaging: The use of volumetric imaging is superior to 2D slices for radiomic studies as there is no need to create information by interpolating or re-slicing.
Multiple sequences: The use of multiple conventional MRI sequences provides additional data and more useful information for radiomic studies.
Multiparametric data: The use of multiparametric data for example diffusion,
perfusion and spectroscopy in MRI will greatly enhance the level of data and could be revolutionary in combination with machine learning,6 given that multiparametric MRI provides information about the tissue microenvironment.
Number of studies: Generally,
at least 100 patients are required in radiomic studies,
but larger numbers will give more power.2 If there is heterogeneity in the dataset,
from the use of different scanners,
different protocols or scans from different institutions,
larger numbers will be required to compensate for the variation.
Pseudonymisation/Anonymisation
In order to comply with research governance guidelines,
imaging studies should be pseudonymised or anonymised prior to analysis.
This is possible through research PACS environments,
supplied by a number of vendors.
Identifiable data is removed and a linking code generated to allow correlation of imaging with clinical data.
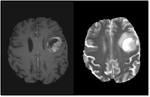
Image Pre-processing and Co-registration
Pre-processing of images involves ensuring that the images are free from artefact,
inhomogeneity correction,
intensity normalisation and re-slicing if required.7 Multiple MRI sequences also require co-registration to allow for segmentation and radiomic feature extraction ( Fig. 1).
Lesion Segmentation
Segmentation of the lesion,
usually a tumour,
is required prior to running radiomic feature extraction.
This can be manual,
semi-automatic or automatic.
There are a number of tools available,
however it is worth noting that manual segmentation of tumours is normal practice for example in radiotherapy planning.
The manual technique can be accurate but is subjective and time-consuming,
particularly with the combination of big data in radiomic studies.
Semi-automatic methods are preferred and involve automatic computer-aided segmentation methods,
most commonly edge-based detection methods in combination with a manually placed seed point,
followed by manual adjustment.
Fully automatic methods can be used,
although these methods may require multiple MRI sequences and may not be accurate where there are complex lesions.
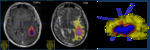
Volumes of interest are created (Fig. 2) which can be loaded into radiomic packages for feature extraction.
Radiomic Feature Extraction
Radiomic feature extraction is performed using dedicated software packages or custom built applications for specific features.
Features consist of shape and size,
first-order statistics (histogram-based techniques of voxel values),
second-order statistics (textural features defining correlations between voxels) and higher-order methods (such as the use of filter grids).7 Examples of textural second-order features include: Grey Level Co-occurrence Matrix (GLCM),
Grey Level Run Length Matrix (GLRLM),
Neighbouring Grey Tone Difference Matrix (NGTDM),
and Grey Level Size-Zone Matrix (GLSZM) features.
A large number of features are extracted; the most significant features are therefore selected using statistical methods before proceeding to machine learning analysis.
Machine Learning Analysis
The selected highly significant features undergo classification (machine learning analysis) along with clinical data and/or genomic data,
to create prediction models using features from the radiological images.
There are a number of different classifiers that can be used,
however the most popular currently include support vector machine (SVM) and logistic regression classifiers.7 The ground truth (reference standard) which is being correlated with radiomic features will be clinical outcome data,
radiology reports,
genomic data or histopathology (Fig. 3).
It is important to ensure accuracy of this data and understand limitations of this information (for example error rates in histopathology) before use as ground truth.
Testing and Validation
Radiomics and radiogenomics prediction models should be validated with an independent dataset,
separate to the training dataset.
Ideally this should be from a different institution.8 As there are many options and methods for lesion segmentation,
radiomic feature extraction and machine learning analysis,
with no standard approach and lack of protocols,
it is therefore very important to ensure robust validation of radiomic and radiogenomic prediction models.
After potential radiomic imaging biomarkers have been identified,
radiologists should attempt to correlate with other conventional imaging features and physiological/metabolic information from multiparametric MRI,
and explain the findings with biological features of disease.
This is an important step in the process of radiomic biomarker validation.