This review appraises studies which use machine learning algorithms in order to generate prognostic information from medical images.
The field of radiomics involves extracting and analysing quantitative imaging features which can then be correlated with other clinical and pathological parameters in order to generate predictive and diagnostic models of disease.
A traditional model of radiomic analysis,
as described by Kumar et al.(2012),
typically involves the following steps: image acquisition,
image segmentation,
feature extraction,
feature selection,
and informatic analyses (1).
The majority of papers appraised in this review utilised a traditional radiomic approach to generate a range of features from the tumour and peritumoral areas and then applied machine learning classification algorithms (most commonly support vector machine and random forest classifiers) in order to correlate these extracted imaging features with various pathological or genetic parameters to identify imaging biomarkers of prognostic and diagnostic interest which are not visible with the human eye.
A critical step prior to radiomic analysis is the process of image segmentation.
Segmentation can be done manually (i.e.
physically drawing around tumour),
or can be automated,
or can be a hybrid of the two techniques (i.e.
manual checking and editing of automated segmentation).
After the area is selected,
individual features (like lesion size,
shape,
heterogeneity) can be extracted.
The imaging data is then analysed by an appropriate machine learning algorithm classifier and the strongest associations between the quantitative imaging features and the chosen clinical,
pathological or genetic outcomes are identified.
Several outcome measures were assessed in the analysed studies such as overall survival,
degree of tumour infiltration,
and genetic mutational status.
The next phase of the process involves testing the predictive model in order to confirm its validity and potential clinical usefulness.
Testing can be carried out on the training data set itself (in the form of “leave one out” cross-validation),
and/or on a separate test data set of imaging that has not been previously seen by the machine.
Imaging data
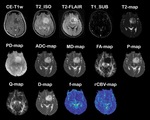
10 of the 18 studies used multiparametric MRI data for image analysis.
For example,
Kazerooni et al,
2018 (2) used T1,
T2,
T1 with contrast,
T2 relaxometry,
diffusion weighted imaging,
diffusion traction imaging,
and dynamic susceptibility contrast imaging as part of their analysis.
While other studies,
like the two papers published by Li et al in 2018 (3,4),
only used T2 MRI sequences.
While too many imaging inputs can increase noise,
too few inputs limit the diagnostic accuracy.
Even with multiple sequences available,
not all studies used all sequences in their analysis.
For example,
Upadhaya et al,
2015 (5) had access to T1 pre contrast,
T1 post contrast,
T2,
and FLAIR sequences,
but only analysed two sequences at a time.
While most studies used MR imaging,
the Imani et al,
2014 (6) paper used FDG PET as part of the input data.
As pointed out by Zhou et al,
2017 (7),
limiting the input data to only imaging,
as was the case with the majority of papers on this topic,
omits a lot of relevant information from the analysis.
Just as a radiologist uses ancillary clinical information when interpreting imaging,
machine learning programs should also have such information available to them when generating algorithms. However,
this was not universally the case,
with Chang et al.
2016 (8) developing a machine learning model including both clinical data and quantitative tumour features to predict progression-free survival and overall survival in patients with recurrent glioblastoma treated with bevacizumab.
Image segmentation
While the majority of studies appraised in this review utilised machine learning for the role of classifying the importance and relevance of extracted features as potential clinically important prognostic or diagnostic biomarkers,
a few of the studies (9,10) capitalised on the recent advances in computational processing power and developments in image processing using deep learning and multi-layered convolutional neural networks to enable faster,
more accurate and reproducible image segmentation – a critical step in the radiomic analytic process.
Convolutional neural networks have previously been demonstrated to be highly effective in the segmentation of multi-modal medical images,
and the immense potential in diagnostic imaging is still to be realised (11).
Li et al,
2017 (9) unified the radiomic analytic process by using deep learning with a 6-layered convolutional neural network in order to both automatically segment the lesion and also extract and select features from the final layer,
for correlation with isocitrate dehydrogenase 1 mutational status.
This combined process of automated segmentation and feature extraction solves several problems,
including the inconsistency and lack of reproducibility using manual or semi-automatic segmentation methods,
whilst also increasing the speed and precision of tumour segmentation.
The range of outcomes studied in this series of papers reflect the diversity of clinically useful biomarkers that are currently being investigated,
and evidences the breadth of potential machine learning has.
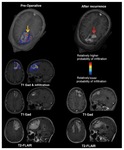
Akbari et al.
2016 (12) found machine learning to be 91% sensitive and 93% specific in its prediction of early glioma recurrence.
The Chang et al.
2016 (8) study found a statistically significant predictive capacity for overall survival in both the training and testing cohorts. Wiestler et al.
2016 (13) created an algorithm that predicted WHO grade status accurately in 92%.
The Imani et al.
2014 (6) paper devised an algorithm for the differentiation of brain tumour progression from radiation induced necrosis,
which was 83% accurate.
Other studies created algorithms able to predict IDH (10),
p53 (3),
ATRX (4),
1p/19q codeletion mutational status (14).