Keywords:
Not applicable, Observational, Artifacts, Computer Applications-General, CT, eHealth, Computer applications, Artificial Intelligence, Artificial Intelligence and Machine Learning
Authors:
M. Patwari1, R. Gutjahr2, R. Raupach3, A. Maier1; 1Erlangen/DE, 2Nürnberg, Deutschland/DE, 3Forchheim/DE
DOI:
10.26044/ecr2020/C-03463
Methods and materials
Data: Ten CT body scans were reconstructed using weighted filtered backprojection [13]. Each volume was reconstructed with the standard clinical dose and with doses of 50%, 25%, 10% and 5% of the standard clinical dose to simulate reduced dose acquisitions. The images reconstructed with 100% of the standard dose were treated as ground truth images. Each image had a resolution of 256 x 256.
The testing dataset consisted of three body CT scans which were not present in the training dataset. The test dataset was reconstructed at standard dose and at reduced (25%) dose.
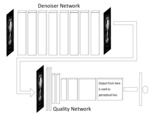
Methods: An eight layer residual CNN [14] was used as our denoising network (Fig. 1). The network contained 4 convolutional layers and 4 deconvolutional layers. There are two skip connections in the network. Each layer contained 96 filters. The filters were of size 3 x 3 for the 2D networks, and of size 3 x 3 x 3 for the 3D networks. 2D and 3D variants of the denoising networks were trained to observe if the inclusion of neighbouring slice information was helpful for denoising.
The network was trained with mean squared error as well as with the use of a deep learned perceptual quality network (Fig. 2) as optimization targets. The output from the 8th convolutional layer of the perceptual quality network was used as a secondary optimization loss (Fig. 3). The training was performed over 10,000 iterations. The updates were calculated using backpropogation and applied using Adam optimiser [15] with a learning rate of 10-4.
Metrics: PSNR was used to measure the noise reduction performance of the network. SSIM was used to measure the feature preservation after denoising.

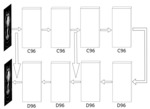
![Fig. 2: This image shows the reconstruction quality network used to extract features. The network has 8 convolutional layers and one fully connected layer. It predicts an IRQM [12] score.](https://epos.myesr.org/posterimage/esr/ecr2020/156576/media/846307?maxheight=150&maxwidth=150)