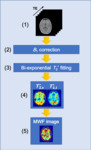
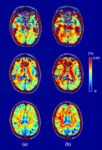
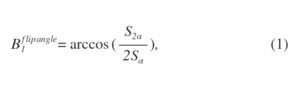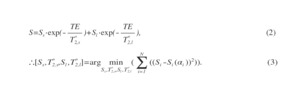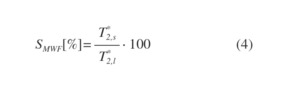Fig. 1 shows the procedure for producing a MWF image. On a 3.0 Tesla MR scanner system (Discovery MR750, GE Healthcare, Waukesha, WI, USA) and a standard head coil receiver, a healthy volunteer (24-year-old male) study was performed using three-dimensional spoiled gradient-echo (3D-SPGR) with and without FS. The imaging parameters were as follows: TE, 0.032, 2.2, 4.6, 6.9, and 10 ms; repetition time (TR), 265.1 ms; field of view, 200 mm; matrix size, 256×256; band width, 976.6 kHz; slices, 26; slice thickness, 5 mm; flip angle (FA), 24 degrees. Then, the TEs were set coherently at an in-phase time taking into consideration the signal attenuation effect (i.e., Dixon effect) due to fat phase variation. The CHESS method was used for fat suppression. Next, B1 inhomogeneity correction was performed to the acquired UTE-SPGR dataset. B1 correction was derived from the double angle method. This method uses the MR images of two FAs using a gradient echo (GRE) sequence. The dataset for B1 correction was acquired with 2000 ms TR, 5.8 ms TE, 40 degrees FA α, and 80 degrees FA 2α. The following formula is used for B1 correction:
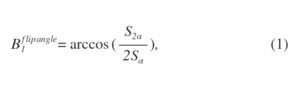
Fig. 2: B1 correction.
where Sα and S2α are the signal intensities from the α and 2α FA data, respectively. Then, calculated B1 was applied to pixel-by-pixel of MR imaging data. Next, T2* values of two-components (T2,s* and T2,l*) were calculated from each UTE-SPGR dataset with and without FS using the bi-exponential fitting model, and then each T2* map was created. Then, the Levenberg-Marquardt algorism was applied. The following bi-exponential fitting model is used [5]:
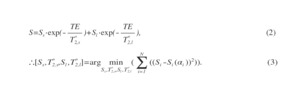
Fig. 3: Bi-exponential fitting model.
Finally, MWF image was generated. The formula for calculating MWF is as follows:
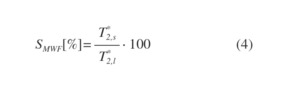
Fig. 4: MWF.
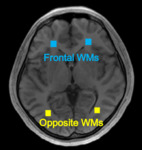
Fig. 5 shows a magnitude image for the Region of Interest (ROI) setting. ROI analysis was performed on different WM areas (bilateral frontal and opposite) of each T2* map and the mean and standard deviation (SD) were measured. The ROI was set to 5 × 5 pixels. The ROI placement is shown in Fig. 5. All imaging analysis was applied using an in-house MATLAB program (Mathworks, Natick, MA, USA).
The results of this experiment are as follows: Fig. 6 shows magnitude images without FS and acquired using multi-echo SPGR.
Table 1 shows each measurement T2* values of frontal WM and opposite WM areas obtained by ROI analysis. T2,s* and T2,l* values with FS showed a lower value than those without FS.
Table 1 Measurement T2*values at each ROI.
|
Areas of ROI
|
With FS
|
Without FS
|
|
T2,s*[ms]
|
T2,l*[ms]
|
T2,s*[ms]
|
T2,l*[ms]
|
|
Frontal WM
|
0.016 ± 0.002
|
44.47 ± 7.53
|
0.019 ± 0.002
|
57.02 ± 7.89
|
|
Opposite WM
|
0.019 ± 0.001
|
72.20 ± 8.12
|
0.021 ± 0.0001
|
73.36 ± 6.09
|
Notes: Data are presented as mean ± SD.
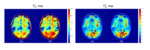
Fig. 7 shows T2,s* and T2,l* maps derived from UTE-SPGR dataset. T2,s* map shows a high signal in the WM area (Fig. 7a, b). T2,s* map without FS showed a slightly higher value than with FS (Fig. 7b, black arrows). On the contrary, T2,l* map showed a low signal in the WM area and there was too great of a difference between the T2,s* maps with and without FS (Fig. 7c, d).
Fig. 8 shows MWF images with and without FS. MWF image without FS shows a higher signal in the WM containing myelin components than with FS (Fig. 8a). As a result, it is possible that axon-derived signals were extracted by applying FS.