Keywords:
Contrast agent-intravenous, CT-Quantitative, Cardiac, Ischaemia / Infarction
Authors:
T. Rienmüller1, C. Baumgartner1, V. Makarenko2, L. Bockeria2, P. Ourednicek3, R. Rienmüller4; 1Hall in Tirol/AT, 2Moscow/RU, 3Brno/CZ, 4Graz/AT
Methods and Materials
Phantom Study
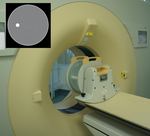
We simulated dynamic CT-based myocardial perfusion measurements using a phantom body containing sterilized water and a small teflon structure (see Fig.
2).
For all experiments 11 slices were acquired over 30 timesteps with a multi-row CT scanner (100kVp/100mAs,
reconstructed slice-thickness of 5mm,
exposure time 180ms,
12Bit).
Image Reconstruction
The images were reconstructed using
- filtered back projection (FBP)
- three different levels (strength) of an iterative reconstruction (IR) algorithm (iDose4,
level 2,
level 4 and level 6).
Image reconstruction was performed applying three different convolution kernels:
- Cardiac Smooth (CA)
- Cardiac Sharp (CC)
- Cardiac Detailed (CD)
This results in a total of 12 different reconstruction modalities and 3960 images.
Evaluation
- The water region was segmented and mean value,
standard deviation (SD) and minimal (MIN) values were determined (Maximal values were not determined,
since the were at the same level due to the transition to the teflon structure.)
- A smaller region of interest (ROI) in the water area was selected (69 voxel) and mean value,
standard deviation (SD) and MIN as well as MAX values were determined.
- A ROI was placed in the part of the image representing the teflon structure.
Again,
mean,
SD,
MIN and MAX were determined.
- Different parameters proposed in the literature (see e.g.
(Jang,
et al.
2011)) for quantitative assessment of image quality were computed: mean square error (MSE),
peak signal to noise ratio (PSNR),
normalized absolute error (NAE) with an idealized image of the phantom as a reference.
Perfusion Measurement
In the next step,
dynamic CT-data of one patient were reconstructed applying FBP and two different levels of the IR algorithm (level 3 and level 7) with a CA reconstruction kernel.
The images (13 slices,
40 time steps) were acquired after contrast agent administration using the same multi-row CT scanner as in the phantom study (100 kV,
100 mAs,
reconstructed slice thickness of 5 mm,
exposure time 180 ms).
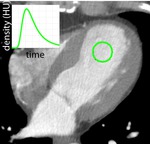
A ROI was placed in the left ventricle and the HU-value-time curves registered (see Fig.
3).
The left ventricular myocardial tissue was segmented semi-automatically.
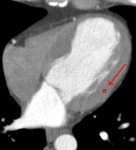
In addition,
smaller ROI (64 voxel were selected by chance) was placed manually in the myocardium in order to compute local myocardial perfusion values (see Fig.
4).
Evaluation
Statistical values of myocardial tissue were computed (mean,
SD,
MIN,
MAX)
for all slices of myocardial tissue and the smaller ROIs.
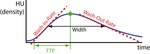
MPF and several conventional parameters describing MBF,
such as Time-To-Peak (TTP),
Wash-in-Rate,
Width and Area-Under-Curve (see Fig.
5),
were compared for all different reconstruction methods.