Purpose
To evaluate the correlation between CT-derived quantitative texture features,
histologic subtypes (WHO) and staging (Masaoka,
TNM) in patients with thymic neoplasms[1].
Methods & Materials
In this retrospective study,
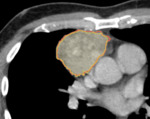
CT scans of patients with histologically confirmed and categorized thymic neoplasms were extracted from the PACS and reformatted to a fixed slice thickness of 2 mm and an in-plane resolution of 1 x 1 mm².
Tumors were manually segmented using the open source software 3D Slicer[2].
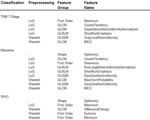
Either using the original image or applying filters (Laplace of Gaussian filter,
Wavelet filter),
pre-defined,
3D-based texture features of 7 major categories were calculated (First Order Statistics,
Shape-based Features,
Gray Level Cooccurence Matrix,
Gray Level...
Results
Patient Statistics
CT scans of 15 Patients (4 female patients,
mean age 51±12 years,
range 30-69 years) could be included for this preliminary analysis.
Subcategories of thymic neoplasms were distributed as follows: WHO A: 2,
AB: 3,
B: 7,
C: 3; Masaoka: I: 2,
II: 6,
III: 3,
IV: 4; TNM T-Stage: I: 8,
II: 0,
III: 2,
IV: 5.
Feature Selection
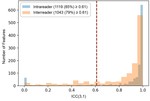
Of the initial 1316 features,
352 were excluded due to poor to moderate intra- or interreader agreement (Figure 2).
Correlation-based attribute selection resulted...
Conclusion
Preliminary results indicate that machine learning-based classification based on quantitative,
CT-derived texture features may be able to correctly predict thymic neoplasm subtypes.
This system could potentially be used to create radiomic signatures as biomarkers.
References
[1] Rashid,
O.
et al.
(2013) Thymic neoplasm: a rare disease with a complex clinical presentation.
J Thorac Dis.,
5(2): 173-183
[2] Kikinis,
R.
et al.
(2014) 3D Slicer: a platform for subject-specific image analysis,
visualization,
and clinical support. Intraoperative Imaging Image-Guided Therapy,
Ferenc A.
Jolesz,
Editor 3(19):277–289 ISBN: 978-1-4614-7656-6 (Print) 978-1-4614-7657-3 (Online)
[3] van Griehuysen,
J.J.M.
et al (2017).
Computational Radiomics System to Decode the Radiographic Phenotype.
Cancer Research,
77 (21),
e104-e107
[4] Shrout PE,
Fleiss JL.
Intraclass correlations: uses in assessing rater reliability....
Personal Information
Christian Blüthgen,
M.Sc.
Institute for Diagnostic und Interventional Radiology
University Hospital Zurich
Rämistrasse 100,
CH-8091 Zurich
E-Mail:
[email protected]