Part One: Support Vector Machine Learning - Liver Fibrosis
In 2012,
we applied support vector machine learning (SVM) to a cohort of 28 patients with hepatitis C to assess textural differences in MRI images [2] to build a predictive model for categorising the patients’ Ishak fibrosis scores [3].
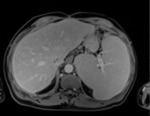
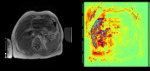
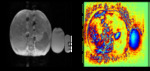
Figure 1 shows an example liver image used in the study.
The assumptions and limitations of the modelling of small patient numbers,
that include cross-validation and overfitting,
are presented.
The small subject population was divided into both training sets and validation sets using a “leave one out" cross validation technique.
Features in the images associated with liver fibrosis (as assessed by biopsy) were identified using clustering techniques.
The parameter optimisation was performed using 27 of the 28 patient image sets as the training data set and then the performance of the resulting model was tested on the 28th patient as the test set.
This was repeated 28 times,
each time leaving out one of the image sets as the single test case.
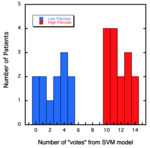
The SVM model yields a number of “votes” for whether or not the image has features associated with liver fibrosis.
The results from the training model for the 3-minute post contrast 3D VIBE images are shown in Figure 2.
The subjects were re-scanned 6 weeks later.
Using the leave one out models trained on 27 of the 28 visit 1 images sets; 27,
28,
and 28 out of 28 cases were correctly classified in visit 1 data for pre-,
3-mins post,
and 10 mins post-contrast images,
respectively (Table 1).
The model trained from all 28 visit 1 datasets correctly classified 23 out of 27,
and 26 out of 28,
and 25 out of 28 cases for the pre-,
3-mins post,
and 10 mins post-contrast images from visit 2 (Table 2).
Table 1.
Summary of the results for the leave-one-out classification of visit 1 MRI data
| Image Type |
Classification Success Rate |
| Pre-contrast |
27 out of 28 cases (96%) |
| 3-min post contrast |
28 out of 28 cases (100%) |
| 10-min post contrast |
28 out of 28 cases (100%) |
Table 2.
Summary of the results for the classification of visit 2 MRI data based on the model trained on visit 1 data.
| Image Type |
Sensitivity |
Specificity |
| Pre-contrast |
80% |
92% |
| 3-min post contrast |
93% |
92% |
| 10-min post contrast |
93% |
85% |
While these results are promising,
the validation at visit 2 was on the same patients involved in the training of the SVM model.
There is a danger of “overfitting” the model,
especially when a small number of cases is involved.
Further studies on independent cohorts of patients are now required.
|
Cross validation assesses how the results of a statistical analysis will generalise to an independent data set.
It is mainly used where the goal is to test the model’s ability to predict new data that were not used in training it,
and to give insight on how the model will generalise to an independent dataset.
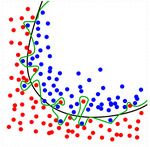
Overfitting is the production of a prediction model that corresponds too closely or exactly to the data upon which it is trained and may therefore fail to accurately predict future observations reliably.
See Figure 3.
|
Part Two: Transfer Learning
In 2016,
our focus shifted from attempting to predict the pattern of liver fibrosis to predicting liver iron concentration (LIC) from MR images.
Our rationale was that we needed to work on problems where we had larger datasets available to us to assess the influence of the number of datasets on the machine learning and to have sufficiently large and diverse populations for validation of the models.
We had available to us several hundred sets of MR liver images acquired using the FerriScan® protocol together with associated liver iron concentrations.
The FerriScan method involves acquiring 11 abdominal axial slices through the liver.
Analysts choose the most appropriate slice and manually segment the liver to carry out the liver iron concentration analysis.
To model the relationship between the images and the liver iron concentrations,
transfer learning techniques with a convolutional neural network (CNN) were used [4].
CNNs have been found to be effective at building models from images [5] and this method along with gradient boosting machines (GBMs) was chosen for this project.
GBMs are efficient for building models from numerical data [6].
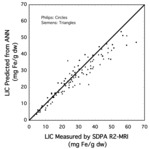
Our initial attempt used 200 MRI datasets to further train a pre-existing artificial neural network that had been trained on 2.6 million human face images of 2600 people [7].
The MRI data comprised images where the image slice for analysis was chosen manually to match exactly the image slice that had been used in the FerriScan® analysis.
By splitting the 200 datasets into training and validation sets,
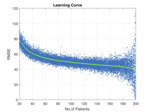
we were able to determine how the number of datasets in the training set determined the accuracy of the model (as determined by the magnitude of the square root of the mean squared difference between the model prediction and the liver iron concentration measured by FerriScan® - see Figure 4).
It can be seen that the accuracy of the model improves (RMSE decreases) as the number of images in the training set increases.
This is to be expected – more data means better models.
As the number of patients approaches 199 the spread about the line increases.
This is because the number of patients in the test set is decreasing (at 199 there is only one patient in the test set) so the measurement error increases.
From this learning curve,
we decided to increase the number of training images to 400 based on the concept of diminishing returns.
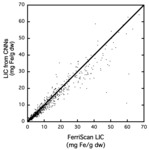
The resulting model was then tested on 100 subjects not involved with the training of the CNN and using scanners different from those used to train the CNN [8].
The results of the testing are shown in Figure 5.
The advantages of a machine learning approach over a manual approach using a parametric formula (such as FerriScan®) are potentially that:
- the machine can use other organs in the image set besides the liver to do its calculations;
- it can use all slices in the image set instead of just the best slice;
- it can figure out and allow for scanner differences;
- it can figure out and allow for patient differences such as age,
gender etc;
- it can learn the patterns of and account for noise in the images;
- it can work out its own formula for LIC instead of being restricted to the bi-exponential formula.
|
CNNs are suited to regression and classification problems which produce a prediction model,
typically of decision trees,
built in a staged fashion to allow optimisation of the mathematical function.
For the CNNs the MatConvNet Toolbox [9] for MATLAB was chosen and for GBM the xgboost package [10] for R was used.
There are two ways to use transfer learning with CNNs.
One way is to leave some of the CNN layers fixed and to retrain other layers to model the MRI images.
The other way is to extract “activations” from one of the CNN layers and to feed them into a separate machine learning model such as GBM.
The former method is problematic because so many parameters are retrained in the CNN that with only a few hundred images it is possible to model the images perfectly and predict LIC exactly.
This model would be overfitted which means that it is specific to images used in the training and would not generalise to unseen images.
GBMs have fewer parameters and so are less prone to overfitting.
They also allow the use of extra information such as age and gender of patients,
although this was not used for the initial proof of concept.
|
|
In our example,
the CNN trained on images other than MRI images.
We used the vgg_face CNN descriptor from Oxford University [6].
This network was trained on 2.6 million face images of 2600 people.
We tried other pre-trained CNNs and this was found to be the best.
The reason may be due to the similarity in shape and features between faces and MRI abdominal images.
This descriptor requires colour images,
so we colourised the MRI images by creating a gradient of skin tones to map the grey pixels onto.
Skin tones were scraped from face images on the internet.
Some trial and error was used to find the best activation layer and we found this to be the fully convolutional layer “fc6.” This layer has 4096 numerical values.
So in essence the vgg face CNN was used to transform a 256 x 256 pixel MRI image into 4096 numbers in a way that retained the essence of the image.
In addition,
we used the 65536 raw pixel values as inputs to a GBM to find which pixels were most valuable in predicting LIC.
The most valuable pixels were,
as to be expected,
clustered around the liver.
We also found that pixels near the edges were valuable – possibly these had something to do with calibration of the MRI images.
We chose the 4096 most valuable pixels and the 4096 activations to create a GBM model that predicted R2 values from the 8192 inputs.
We used 5-fold cross validation to prevent overfitting for all our models.
|
Limitations of the approach:
- the number of samples provided for training were small;
- the low sample count forced the choice of transfer learning to attempt to solve the problem;
- manual selection of image slice and manual segmentation of the torso were required.
Part Three: Deep Learning and Convolutional Neural Networks
In 2017,
we were able to access a much larger dataset of MR images (several thousand) with associated liver iron concentration values.
With the much larger dataset,
we decided to approach the problem by
- training CNNs from scratch rather than using transfer learning;
- training a CNN to select the image slice for analysis;
- training a CNN to predict the liver iron concentration without the need for manual image segmentation.
The new approach leveraged the latest open source deep learning libraries (see box below for details).
For training image slice selection,
the original slice number chosen manually by FerriScan® analysts was used to train a CNN.
For training liver iron concentration prediction,
the resulting FerriScan® result was used to train a separate CNN.
|
The machine used contains a single 1080ti NVIDIA graphics card,
128 gigabyte of memory and a 1 terabyte NVMe ssd drive.
The open source deep learning libraries used were Tensorflow & Keras.
|
After training on 15,354 image datasets,
the trained CNNs were tested on a further 906 new datasets that had not been used in the training of the CNNs.
Furthermore,
the unseen 906 datasets used for testing the performance of the CNNs had been acquired from 77 scanners that had not been used for acquiring the training data.
The predictive performance of the trained CNNs can be seen in Figure 6.
|
Saliency Maps
A question that inevitably arises when using artificial intelligence to analyse images is “what is the CNN doing?” This question is almost impossible to answer but some clues can be given by examining “saliency maps” that indicate the weighting that each pixel in an image has with respect to particular aspects of the neural network that result in the final output.
In our example of the measurement of liver iron concentration from MR images,
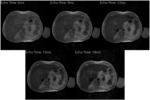
manual analytical analysis involves modelling the exponential decay of signal in a series of spin echo images (see Figure 7). The rate of decay is related to the liver iron concentration by a calibration curve.
For some scanners,
signal gain changes significantly between echoes and hence analysts normalise tissue signal intensities to that observed in the normal saline bag that is scanned along side the patient. Analysts also use the signal variation in subcutaneous fat to help model the RF signal intensity throughout the torso. With these observations in mind,
it is very noteworthy that saliency maps generated by the CNN used for liver iron concentration measurement show that significant attention is paid to the liver (as expected),
but also to the subcutaneous fat (Figure 8) and also to the normal saline bag (Figure 9) for scanners that change gain between echoes. The startling aspect of these saliency maps is that they show that the CNN has “learned” to examine these relevant parts of the images without having been explicitly trained or instructed to do so!
|
Lessons Learned
- Thousands of datasets were required to train the CNN from scratch;
- Reasonable results can be achieved with large numbers of well labelled data;
- CNNs can completely automate an analysis process.