Keywords:
Artificial Intelligence, Neuroradiology brain, MR, Contrast agent-other, Cancer
Authors:
S. Bhardwaj1, C. Chong1, M. Agzarian1, S. Pati2, U. Baid2, S. Bakas2, M.-S. To1; 1Bedford Park, SA/AU, 2Pennsylvania, Philadelphia/US
DOI:
10.26044/ranzcr2021/R-0362
Results
In a separate study, the initial phase of the federation was conducted with 11 collaborating sites, and it focused on addressing any potential network communication issues and with the segmentation task being focused solely on the tumor core region. The conclusion of the initial phase yielded an improvement in the DICE Similarity Coefficient for the tumor core of 11%, when comparing the performance of the consensus model with the model trained only with publicly available data. By conducting this study with much larger number of collaborating sites, we expect that this improvement in all segmented regions will be much larger when the consensus model of the complete federation is evaluated.
We describe the utilization of the FeTS platform, which is a user-friendly tool allowing for collaboration of institutions across the globe to train a consensus DL model for medical imaging without sharing patient data. The consensus model, derived by aggregating training results from multiple collaborating sites, underwent multiple iterations of training until convergence.
Stage 1. Data pre-processing.
The T1, T1ce, T2 and FLAIR images were pre-processed using the Brain Tumor Segmentation (BraTS) pipeline, made publicly available through the FeTS tool and CaPTk [2].
Stage 2. Preliminary segmentation and manual correction.
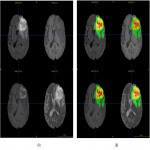
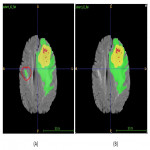
An initial inference step using the label fusion of pre-trained segmentation models was performed. The preliminary segmentations were individually reviewed and manually corrected. The most common errors of the preliminary segmentation included:
- Incorrect segmentation of hyperintensities white matter non-glioblastoma lesions as tumor lesions (Figure 2).
- Incorrect segmentation of choroid plexus and blood vessels within the brain parenchyma ET regions.
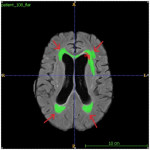
- Incorrect segmentation of periventricular areas edematous tissue (Figure 3).
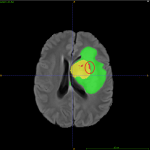
- Necrotic core of the tumor was incorrectly labelled outside the ET region (Figure 4).
The most incorrectly segmented brain tumor region was NET. We found that the tumor core (NET) boundaries were not clearly delineated and were often confused with ET or ED regions by the preliminary model (Figure 4).
Stage 3. Federated learning and consensus model.
Federated learning was performed with the corrected segmentations, generating the final consensus model whose performance will be evaluated in all segmented regions.