Aims and objectives
The development of machine learning and AI systems in healthcare critically relies on large well-labelled datasets with adequate data volume,
annotation,
truth,
and reusability [1].
Unfortunately,
it has been cited that the day-to-day radiologist error rate is on average 3–5% and in medicine the rate of missed,
incorrect,
or delayed diagnoses has been reported as high as 10-15% [2-3].
As well,
Sadigh et al.
found that imaging reports were categorized into either mislabeled or misidentified patient or wrong dictation or report events at a rate...
Methods and materials
Study Population
A retrospective case-control review was performed to export multiple consecutive CT abdomen examinations over a 10-year period at two academic tertiary care hospitals; in order to maximize diversity and dataset variability.
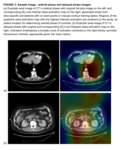
A random sample of images were divided into 6-classes (Fig. 1).
These classes were [A] enhanced slices including liver during the arterial phase [B] slices not including liver during any phase of enhancement,
[C] enhanced slices including liver during the delayed phase,
[D] non-axial slices,
[E] non enhanced slices including liver,
and...
Results
17,465 CT slices were collected from 2 sites between 2013-2017.
The model obtained an overall accuracy of 0.913 on the test set.
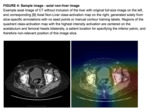
Individual class AUC’s ranged from 0.985 for [C] Liver-Delay to 0.998 for [A] Liver-Arterial (Fig. 2).
All reported accuracies were excellent.
Average training time per epoch was 90 seconds on a single Titan X Pascal (Nvidia,
Santa Clara,
California).
Average time to classification inference per CT Slice was under 3 seconds.
Average time to the generation of the heat map visualizations (i.e.
Salience...
Conclusion
Study Limitations
Ideal classification machine learning experiments have classes of equal sample size.
With unequal sample sizes,
models may be able to correctly classify images based on overall population probability irrespective of image contents.
It is possible that our model inherently favours classifying scans as non-axial or axial non-liver CT slices,
over delayed or non enhanced axial liver CT slices,
although this was not seen to be a major contaminant in the test set classification characteristics.
In addition,
we anticipate that aggregated over multiple slice...
References
[1] Kohli,
M.,
Summers,
R.
and Geis,
J.
(2017).
Medical Image Data and Datasets in the Era of Machine Learning—Whitepaper from the 2016 C-MIMI Meeting Dataset Session.
Journal of Digital Imaging,
30(4),
pp.392-399.
[2] Brady,
A.
(2016).
Error and discrepancy in radiology: inevitable or avoidable?.
Insights into Imaging,
8(1),
pp.171-182.
[3] Bruno,
M.,
Walker,
E.
and Abujudeh,
H.
(2015).
Understanding and Confronting Our Mistakes: The Epidemiology of Error in Radiology and Strategies for Error Reduction.
RadioGraphics,
35(6),
pp.1668-1676.
[4] Sadigh,
G.,
Loehfelm,
T.,
Applegate,
K....