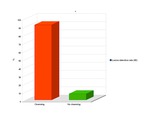
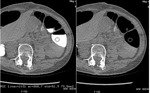
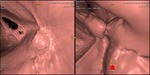
In 2D analysis image quality of cleansed CTC images did not differ significantly from that of source CTC datasets in the various colonic segments (cecum and ascending colon 4.31±0.48 vs 4.54±0.66; transverse colon 4.38±0.65 vs 4.38±0.51; descending colon 3.92±0.49 vs 3.92±0.64,
sigmoid colon 3.38±0.77 vs 3.46±0.66; Wilcoxon signed rank test,
p> 0.05) (Fig.1).
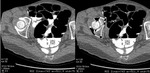
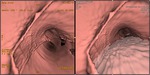
Electronic cleansing allows a significantly higher diagnostic rate concerning 3D images; in fact,
24 findings were correctly identified (92.3%) with cleansing,
than without (7.7%,
Fisher’s exact test p<0.001) (Fig.
2).
Digital colonic cleansing is a prerequisite for 3D post- processing of tagged CTC dataset,
provided that cleansing works in an optimal way; this point seems highly dependent on adequate stool labellig rather than the action of cleansing itself.
In fact,
the persistence of stool makes tagging inhomogeneous and cleansing inefficient,
leading to various types of artifacts that can be noisy in 2D and compromise the interpretation of 3D images.
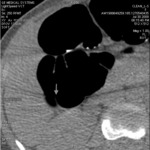
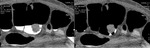
Inhomogeneous tagging can make reading impossible,
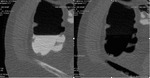
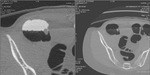
causing anomalous subtraction in extracolonic structures and intraluminal contrast persistence in particular in sigmoid colon; this can occur in case of too early scans (less than 3 hours after oral administration of tagging material) where the density of tagging material can be under the threshold of subtraction (Fig.
3- 4).
Blurring is an edge artifact and results in an ill- defined aspect evident in mucosal- lumen interfaces,
it is related to the inner actions of the various algorithms at the level of the interfaces and can be annoying without causing interpretative mistakes in 2D as well as in 3D,
while tagging remnants may determine interpretative mistakes in 3D (fig.
6- 7).
Electronic cleansing does not lead to a significant improvement of 2D images,
and cleansing-related artifacts although noisy did not impair the diagnostic yield of CTC datasets in any case.
Furthermore,
in 2D interpretation,
no conspicuity,
morphology or size modification of lesions were observed (fig.
8)
This algorithm is also handling in use; for both readers,
the mean evaluation times with subtracted images was approximately 1.5 min longer that with native images in 2D.
The reliability of the cleansing algorithm quantified in 2D have kicked off its evaluation in 3D.
Without cleansing 3D evaluation is not possible,
as demonstrated in our data (fig.
9- 10); with cleansing,
sensitivity is high,
particularly in datasets with homogeneous,
high density tagging.
In case of poor quality of tagging,
images in 2D can be noisy and lesions can be missed in 3D (fig.
11).