For this ethics board approved trial,
11 volunteers (three male and eight female,
aged 20 to 38 years) were recruited,
which were in average training state and did not participate in regular physical training in the period preceding the trial.
Written informed consent was obtained from all participants.
All volunteers were scanned in two sessions: before and after a 6 – 8 week training period (variation depending on scanner availability).
For the exercise program volunteers were assigned to train the same leg,
3-4 times a week.
Each exercise session was composed of 3 to 4 sets of unilateral squats until muscular fatigue was reached.
The number of squats was recorded in an exercise journal.
Physical activity of the other leg was limited to normal daily activation; general physical exercise or sports were prohibited for the study period.
During each scanning session volunteers were scanned twice.
After the initial scan,
bilateral squats were performed until muscular exhaustion; scanning was repeated,
to measure the influence of the single exercise session as well as the influence of the longer training period.
All MRI scanning was performed on a 3 Tesla whole body scanner (Magnetom Verio,
Siemens,
Erlangen,
Germany) in prone position with matrix coils.
Axial slice groups were positioned 20 cm above the knee joint as defined by scout images.
The field of view was 400 x 400 mm,
with a matrix of 384 x 384 yielding an in plane resolution of 1mm with a slice thickness of 5mm.
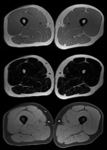
Scanning sequences include a gradient echo sequence with two different echo times for in-phase and opposed-phase images (three-dimensional (3D),
10 slices,
TR 20ms,
TE 2.45 / 3.675 ms,
flip angle 15 deg) to calculate fat- and water-maps (figure 1).
Images were evaluated using an in-house developed evaluation tool (Mathlab R2001a,
The MathWorks Inc.,
Natick MA,
USA).
For muscle selection,
ROIs (Figure 2) were drawn outlining the large muscles of the thigh (rectus femoris,
vastus intermedius,
medialis and lateralis,
semimembranosus,
semitendinosus,
biceps femoris,
sartorius,
gracilis) using free software (ITK-SNAP,
Penn Image Computing and Science Laboratory (University of Pennsylvania),
Philadelphia (PA),
USA) [7].
Mean fat fraction and cross sectional area in the middle slice were calculated and compared between scanning sessions and legs.
Data were compared using the method described by Bland and Altman [8] and differences were calculated with Wilcoxson-rang sum tests using a statistical software package (JMP 9,
SAS Institute,
Cary (NC),
USA).