We developed a client-server software solution that allows a radiologist to record key image findings from any picture archiving and communication system (PACS) or advanced imaging workstation while he or she dictates descriptions of those findings.
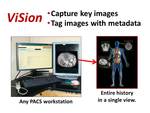
The ViSion client software runs in parallel with any PACS or advanced imaging software that operates on a computer workstation using the Windows® operating system (Figure 2).
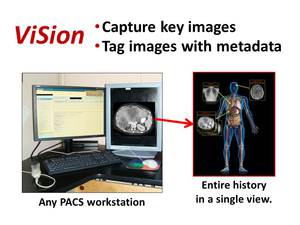
Fig. 2: The ViSion system works by capturing key images from any PACS or advanced imaging workstation, tagging the images with anatomy and pathology terminology extracted from verbal dictations, and assembling a multimedia report on a computer server that is accessible via a web browser.
As a radiologist identifies image findings,
the he or she presses a speech microphone or keyboard button to initiate a screen capture and record a verbal description of each finding.
The image and speech data are uploaded to a cloud-based server where metadata describing the anatomical location,
radiological observation/diagnosis,
disease metrics,
medical priority,
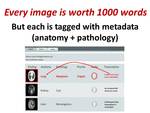
and target lesion designation are extracted from the transcribed speech and used to tag the images in a database (Figure 3).
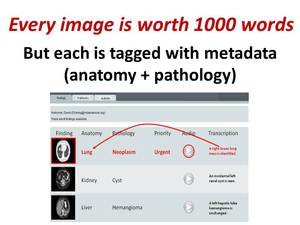
Fig. 3: A picture is worth 1000 words in whatever language is spoken, but ViSion tags each key image with just two words describing a finding's anatomical location and pathology (i.e., radiological observation or diagnosis).
Alternatively,
a radiologist may use pull-down menus in the reporting system to tag images.
The data are then assembled into a multimedia structured report that organizes the image findings by anatomical categories. Alternatively,
ViSion can display the image findings in a graphical representation of a patient with image icons linked to specific anatomical sites.
The assignment of a medical priority to each finding on a 5-point scale (from Incidental to Life-threatening) enables the automatic notification of critical results when a ViSion report is signed by the radiologist.
The medical ontology (controlled vocabulary with defined relationships between terms) used to support ViSion is described in another EPOS poster,
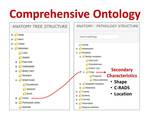
“Development of the ViSion Ontology.” The basic principle of the ViSion ontology is that anatomy and pathology terms are paired to create radiological observations and diagnoses (Figure 4).
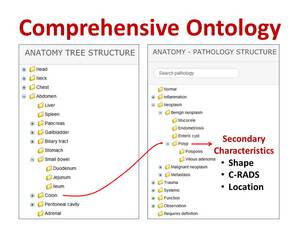
Fig. 4: An observation or diagnosis in the ViSion system is created by the pairing of an anatomy term with a pathology term. The ViSion ontology contains over 12,000 such observations and diagnoses. Secondary characteristics can also be assigned in order to provide additional detail for a particular finding. In this example, the terms "Colon” + “Polyp" are combined, and the secondary characteristics that may be assigned are specific to colonic polyps.
The ViSion ontology contains over 12,000 observations and diagnoses that have been translated to multiple languages.
As a result,
a ViSion report can be automatically translated to any language covered by the system.
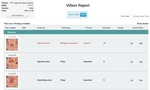
ViSion does not translate the entire narrative dictation associated with each finding but only the metadata which allows for the communication of essential medical information (Figures 5-7).
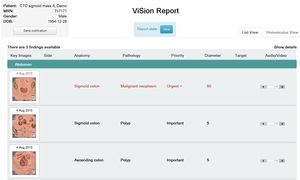
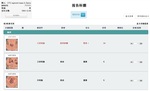
Fig. 5: This is an example of a ViSion report generated in English. In addition to the labeling of an image finding with anatomy and pathology terms, a finding may be labeled with a priority level, disease metric, and target lesion designation. The incorporation of the radiologist's verbal dictation, found under the audio column, allows for the radiologist's unlimited freedom of expression.

Fig. 6: Translation of the ViSion report in Figure 5 to Chinese.
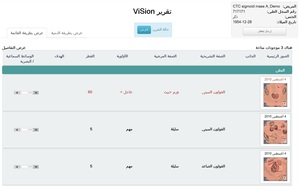
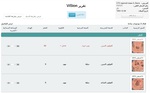
Fig. 7: Translation of the ViSion report in Figure 5 to Arabic.
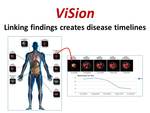
ViSion provides the ability to link image findings from serial radiological examinations in order to generate disease timelines illustrating progression of disease at specific anatomical sites (Figure 8).
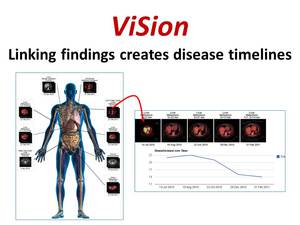
Fig. 8: ViSion provides the ability to link image findings from serial examinations in order to create disease timelines that illustrate the progression of disease at a particular anatomical site with images and graphed metrics.
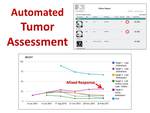
Particular image findings in a ViSion report can be designated by the radiologist as "target lesions" from which ViSion automatically calculates response evaluation criteria in solid tumors (RECIST) curves (Figure 9) (7).
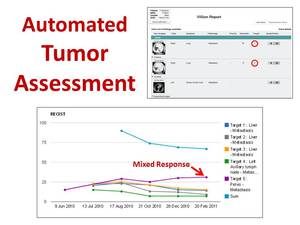
Fig. 9: Specific image findings in a ViSion report can be designated as "target lesions" that are used to calculate RECIST curves. Analysis of individual target lesions may identify mixed response to targeted therapies.
The concept of linking image findings may also be used to associate image data from different medical disciplines,
including pathology,
histology,
and genomics (Figure 10).
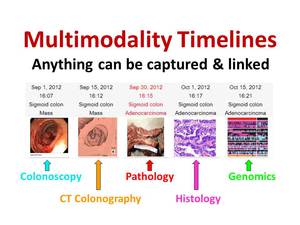
Fig. 10: The linking of image findings is not limited to radiological images but can be applied to multiple image-based medical disciplines. In this example, a colonoscopy image is linked to CT colonography, pathology, histology, and genomic images. Note how the diagnosis changes from a non-specific entity "Mass" to "Adenocarcinoma" as a more specific diagnosis is rendered.
Finally,
ViSion is capable of creating a “composite” report that shows the most recent image findings specific to anatomy,
regardless of modality,
so that the entire history of a patient can be shown in a single display (Figure 11).
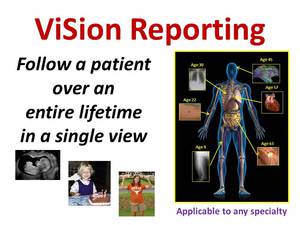
Fig. 11: ViSion's "composite" view shows the entire history of a patient in a single view by displaying the most recent image findings specific to anatomy.