Patient population: A total of 349 chest CT examinations from 324 unique subjects were retrospectively selected from the NHS Lothian database.
Eligibility of CT scans for each group was determined using information from the radiology reports with cross referencing to the electronic health records as appropriate.
Subjects for the first two groups were selected to mimic a lung cancer screening population.
Inclusion criteria were subjects between 50-74 years of age,
current smokers or those with a smoking history and/or reported to have radiological evidence of pulmonary emphysema were found eligible for the first two groups.
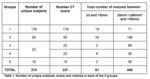
Group 1 consists of 181 CT scans which were clinically reported as being free from pulmonary nodules and group 2 consists of 100 CT scans which were reported to have at least 1 and no more than 10 pulmonary nodule(s).
Group 3 consists of 25 CT scans which were followed up for the presence of a pulmonary nodule,
group 4 consists of the follow-up CT scans of group 3.
Finally,
group 5 consists of 18 CT scans with part-solid and/or ground-glass nodule(s) described in the original radiology report.
Group 5 was intended to increase the overall number of sub-solid nodules.
Specific exclusion criteria were slice thickness >3mm and the presence of diffuse pulmonary disease in the radiology report and/or the CT images,
with widespread abnormalities such as interstitial lung disease,
which is very likely to lead to significant symptoms and therefore didn’t correspond with an asymptomatic screening subject.
Data acquisition: Patients were scanned with Aquilion (n=330),
Aquilion-CX (n=2),
and Aquilion ONE (n=1) CT scanners from Canon Medical Systems (formerly Toshiba Medical Systems),
Otawara,
Japan and LightSpeed (n=2),
LightSpeed Plus (n=2) CT scanners from General Electric Medical Systems,
Waukesha,
United States.
Average tube peak potential was 120 kVp,
(median: 120 kVp,
range: 120-140 kVp).
Average tube current was 243 mAs (median: 232 mAs,
range: 80-491 mAs) and the average CTDIvol was 14.0 mGy (median: 14.8 mGy; range: 2.9-29.7).
Data were reconstructed at a mean slice thickness of 1.0 mm (median: 1.0mm,
range 1.0-2.5mm).
The following reconstruction kernels were used for CT scans from Canon Medical Systems FC03 (n=120),
FC07 (n=99),
FC08 (n=4),
FC10 (n=3),
FC12 (n=7),
FC30 (n=1),
FC51 (n=99) and LUNG (n=3),
STANDARD (n=1) for CT scans from GE Medical Systems.
All CT scans were reconstructed using filtered back-projection.
CAD software: The CAD software evaluated in this study was Veye Chest version 2.0 (Aidence B.V.,
Amsterdam,
the Netherlands).
Image annotation: A two-phase process was developed for the asynchronous interpretation by a panel of three thoracic radiologists with at least 9 years experience in reading Chest CT scans,
JM,
GR and EB,
expert readers 1,
2 and 3,
respectively.
Prior to the start of the study each reader received training on the annotation tasks and how to use the annotation tool.
A comprehensive set of written instructions was available during the entire annotation process.
In summary,
the initial “blinded” phase required readers 1 and 2 to independently perform a free search on all CT scans on a radiology reporting workstation.
In half of the CT scans,
which were selected at random,
the detection results of CAD were made available.
The study design ensured that each CT scan was reviewed twice,
once by each reader,
once by one reader with the results of CAD (AIDED) and once by the other reader without (UNAIDED).
Readers were asked to identify all lesions which they considered to be a pulmonary nodule without clear benign morphological characteristics (i.e.
calcified nodules).
They could mark a pulmonary nodule by adding a manual annotation or classify a CAD prompt as either a true positive or false positive.
They were required to register all nodules that were present on CT scans from both groups 3 and 4,
where possible.
Finally,
the readers also classified all false positive prompts in three different groups: micro-nodules (largest axial diameter <3mm),
masses (largest axial diameter >30mm),
benign nodules (benign calcification pattern or clear benign perifissural appearance),
non-nodules (any finding that could not be classified in any of the other sub-groups).
Subsequently,
non-nodules were further classified as: pleural plaque,
scar tissue,
atelectasis,
fibrosis,
fissure thickening,
pleural fluid,
pleural thickening,
intrapulmonary vessels,
consolidations,
outside of lung tissue,
or other (free format).
After completing all the readings on the workstations the readers subsequently reviewed their own previously identified nodules on a tablet (iPad Pro).
The reader was asked to determine the composition (solid or sub-solid) of the nodule and subsequently segment the nodule on every slice by delineating the border using a stylus (Apple Pencil).
After the blinded phase was completed the results from readers 1 and 2 were evaluated for the presence of any discrepancies.
Discrepancies were defined as a difference between the results in terms of: location (3D dice coefficient of 0); composition; segmentation (3D dice coefficient < -1 standard deviation of the mean 3D dice coefficient) and nodule registration.
The second “unblinded” phase required reader 3 to adjudicate all discrepancies from the blinded phase without the results of CAD,
free search was not allowed.
The review was performed using the same materials used in the blinded phase.
Reader 3 created a third independent reading for each nodule that had a discrepancy for at least one characteristic.
Reference standard: The reference standard for nodule registration was created using CT scans from groups 3 and 4,
subsequently growth rate was determined as the relative volume difference between a nodule visible on a CT scan from group 3 and on a study from group 4.
Data analysis: When looking at nodules visible on sequential scans nodule registration from CAD was scored as either a true positive-pair (TP-pair),
if the detected registration was included in the nodule registration reference standard,
or otherwise as a false positive-pair (FP-pair).
The mean discrepancy between growth percentages determined by readers and CAD alone was calculated.