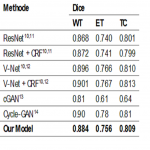
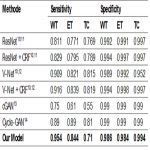
To evaluate the performance of the proposed convolutional neural network (CNN) in this retrospective study, two different test datasets were used and the results of the model on these datasets were compared with some reference segmentation methods that are considered as state-of-the-art in biomedical image segmentation. A CNN model was trained using the training data from these datasets and validated using the test data.
Data:
Whole-Body MRI: MRI scans (acquired with the Dixon technology) of 14 patients without musculo-skeletal tumor disease and without the presence of an external and internal fixator (age 18-90 years). In this work, 3D MRI scans were acquired using the 3T MRI machine with a 3D two-point Dixon VIBE sequence for in-phase, out-of-phase, water and fat imaging. These patients have given their consent for the further use of their data at the time of the examination. Manual segmentation of skeletal muscle and adipose tissue for whole body MRI Dataset was performed by using the ITK-Snap [4] software and validated by an experienced radiologist; they served as the ground truth for training the classifier and performance assessment.
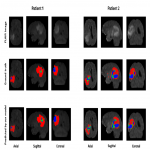
Brain Tumor Benchmark Dataset: For brain tumor segmentation, the BraTS 2018 dataset [5-7]was used. All multimodal BraTS scans were acquired with different clinical protocols and different scanners from multiple (n=19) institutions. Each patient was scanned with FLAIR, T1ce, T1 and T2. Training set consists of 285 cases with ground truth provided. The validation sets and the testing set contain images of 66 and 191 patients with brain tumors, respectively. Since just ground truth for the training set is available, 20% of the training set was randomly selected as the local validation set during training.
Segmentation:
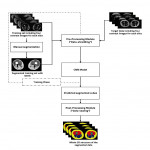
The key principle of the proposed approach is to utilize the image information provided by the four contrast images of the Dixon MRI. The approach can be summarized into three major modules: (1) Pre-processing module (2) The machine learning classifier, is learned from the training set and then used for voxel classification of the target images . (3) Post-processing module.
The overall schematic of the proposed approach is shown in Fig.1.
Pre-Processing: the approach is designed to work with images of any size and dimension, because in the pre-processing phase the data will be "shredded" into several pieces and subsequently fed into our network.
Network Architecture: the architecture is inspired by the 3D-U-Net of Kolařík et. al [8] , which itself is based on the original U-Net implementation [9]. However, for this study, the network was further improved and completed by adding more interconnections between layers.
Post-Processing: in this module, the predicted portions are "sewed" into the original shape of the target dataset and a 3D structure of the segmented data will be generated.