Keywords:
Artificial Intelligence, Computer applications, Musculoskeletal system, MR, Computer Applications-3D, Computer Applications-General, Imaging sequences, Athletic injuries
Authors:
S. Ramedani, H. Von Tengg-Kobligk, C. Morhard, K. Daneshvar Ghorbani
DOI:
10.26044/ecr2022/C-20511
Results
In the first attempt to train the model, the lower limb region was manually segmented with 6 labels, including quadriceps, hamstring group, glutes , bones, subcutaneous fat and intramuscular fat, and trained the model with these labels. The proposed method, achieved good segmentation accuracy in muscle-fat composition, even with a training dataset of only 14 patients, reached an average mean dice coefficient of 0.86 across all classes.
The proposed model is modular and non-specific and can therefore be easily adapted to classification tasks in other domains. To demonstrate its applicability in image segmentation, The model was applied to the BraTS benchmark dataset. Due to hardware limitations, only 10% of all training data used as training set, which includes 28 patinets.
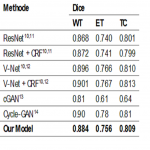
To evaluate the initial experimental results, the average Dice scores, sensitivity, and specificity were calculated for whole tumor (WT), tumor core (TC), and enhancement tumor (ET). Fig. 2 presents the mean values of the dice coefficient of the different models on the BraTS 2018 validation set.
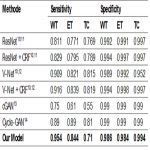
Fig. 3 reports the respective mean values of the sensitivity and specificity.