Retrospective study on consecutive patients with colorectal liver metastases (CRLM) over an 18-month period (July 2006 - December 2007).
To assess tumour burden,
each patient underwent MDCT thorax/abdomen/pelvis,
PET/CT and liver MRI.
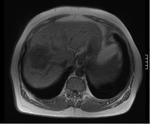
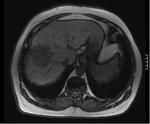
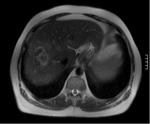
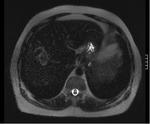
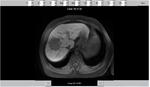
This study looks at the liver MR imaging only which consisted of unenhanced sequences (T1W IP/OP,
T1W 3D GRE + FS,
dual echo T2W,
T2W + FS) (Figures 1 - 5),
dynamic T1W 3D GRE + FS multiphase gadolinium-enhanced sequences at 20s/60s/120s (Figures 6 - 9) and 20 mins delayed T1W 3D GRE + FS gadoxetic acid (Primovist)-enhanced sequences (Figures 10 and 11).
All scans were performed with a 1.5T MRI scanner.
For the purpose of this study,
MRI scans were considered as "standard protocol" (unenhanced and dynamic multiphase post-gadolinium sequences) and "standard + Primovist protocol" (full set of "standard protocol" sequences + 20 mins Primovist-specific sequences).
Primovist is gadoxetic acid (Gd-EOB-DTPA; Bayer Schering Pharma AG,
Germany) 0.025 mmol/kg of 0.25 mmol/mL (10 mL ampoule) (Figure 12).
Patients were included in the study if they had a gadoxetic acid (Primovist)-enhanced MRI and liver resection within 3 months of each other.
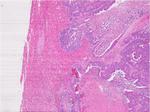
Gold standard: All CRLM were confirmed by pathology and/or intra-operative ultrasound (IOUS) (Figures 13 - 15).
A custom software program was developed to present all images and acquire the rating/location data (Figure 16).
The C++ programming language along with the ITK and FLTK programming libraries were
used.
Readers were first given the "standard protocol" sequences of each of the patients.
After a 2 week interval,
the readers were given the "standard + Primovist protocol" sequences.
Using the software program, 6 readers (2 senior staff abdominal radiologists,
2 newly appointed staff radiologists with North American Fellowship training in abdominal imaging and 2 4th year radiology residents) independently decided whether CRLM or other lesions were present on a per Couinaud segment basis for each patient.
Each liver segment was rated on a 1-6 confidence scale,
with a default value of 1 (Figures 17 and 18).
To maintain uniformity in user interaction,
source PACS windowing and centering were used and "live" image windowing was disabled.
The reader scrolled through the dataset using the mouse-wheel or 2
image-advance buttons (moving 20 images forwards/backwards) at the bottom of the screen (Figure 19).
The image parameters could not be included so the readers were provided with a list of the sequences for each study (Figure 20).
The MRI studies were presented in random order for both readings; there were no identifying features on the studies to link them to a specific patient; readers were blinded to imaging reports,
clinical information and IOUS/pathology reference standard data.
Sensitivity,
specificity,
ROC curves and AUC values were generated using the JRocFit software program [1].
Individual and combined sensitivity,
specificity and AUC values for the readers were compared using DBMMRMC [2].