Keywords:
Computer applications, Oncology, CT, PACS, RIS, Computer Applications-General, Health policy and practice, Structured reporting, Metastases, Neoplasia
Authors:
D. J. Vining1, A. Pitici2, I. Aghenitei2, C. Popovici2, M. Jurca2, R. Rosu2, A. Tsimberidou1; 1Houston, TX/US, 2Chapel Hill, NC/US
DOI:
10.1594/ecr2013/B-0807
Methods and Materials
We initially analyzed the serial radiology reports from ten patients with cancer to determine if radiologists conducted the following (Figure 2):
- Reported tumor metrics
- Saved the measurements in PACS
- Correlated the same tumor metrics consistently across serial examinations

Fig. 2: Analysis of the conventional narrative radiology reports from ten patients enrolled in a Phase I clinical trial. The radiologists reported metrics in 90% of cases, saved those measurements in the PACS in 20% of baseline and 50% of follow up studies, and consistently correlated the same lesions in 40% of patients. (BL exam = Baseline examination; BL Report = metric reported; BL PACS = metric saved in PACS; T1 Report = metric reported for subsequent exam; T1 PACS = metric saved in PACS for subsequent exam; Correlation = consistent correlation of findings between exams)
To address the need for more consistent tumor response assessment,
we created a structured reporting system,
called ViSion,
which is also the subject of another EPOS poster,
“A ViSion for Global Multimedia Structured Reporting.”
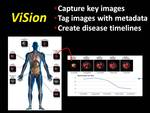
The ViSion system captures and manages image data and information from any PACS or advanced imaging workstation (2).
The client-server software records screen captures of key images plus verbal descriptions dictated by a radiologist,
uploads this data to a computer server where each key image is tagged with anatomy and pathology terminology,
and presents the structured data in a multimedia report (Figure 3).
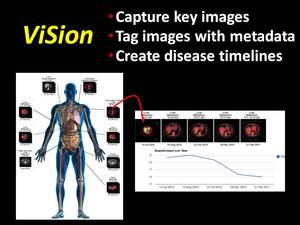
Fig. 3: The ViSion structured reporting system captures key images, tags those images with anatomy and pathology terminology, and presents the data in a unique structured report format. ViSion links image findings across serial examinations and generates disease timelines that illustrate the progression of disease at specific anatomical sites.
In addition,
ViSion provides the ability to link image findings from serial examinations in order to generate disease timelines for each anatomical site of disease.
A disease timeline illustrates the disease progression with both images and graphed metrics.
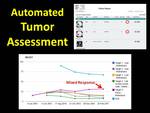
Specific image findings can be designated as “target lesions” (Figure 4).
ViSion sums the diameters of target lesions to generate a RECIST curve illustrating disease progression.
The analysis of individual target lesions may identify mixed response among different metastatic sites which are not evident in the RECIST calculation alone.
The identification of mixed response is important in the era of targeted therapies.
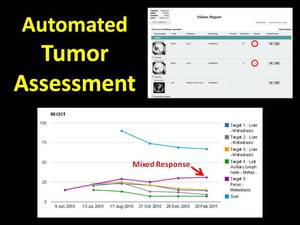
Fig. 4: Specific image findings can be designated as "target lesions" that are used to calculate RECIST curves. Analysis of individual target lesions may indicate mixed response to targeted therapies.