What is radiomics?
Radiomics is the high-throughput extraction and analysis of quantitative imaging features from medical images (1,
8,
9).
There are different groups of features (e.g.,
first-order statistics,
shape- and size-based features,
textural features,
and wavelet features) and each group utilizes a specific mathematical approach to provide information about the image.
Whereas textural features were already used decades ago to analyze chest radiographs (10),
the rapid advances in machine-learning techniques have opened up opportunities in image analysis that surpass previous achievements every year.
It is important to note that the process of radiomics is not simple,
but,
rather,
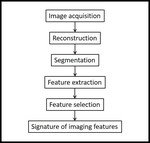
a workflow process that consists of several steps,
each implementing complex computational methods (8) (see Fig.
1).
Radiologists are evaluating images qualitatively every day.
However,
only a small amount of the quantitative information contained in the images can be utilized,
mainly because it is not accessible for intuition- or experienced-based recognition by humans.
Moreover,
many quantitative methods (such as for measuring bone mineral density) are limited to special imaging modalities,
leading to the exclusion of routinely acquired medical images from quantitative analysis.
Providing this kind of information to the radiologist in a comprehensible way would greatly benefit the diagnostic value of a radiologist’s report.
A very similar situation can be found during daily rounds where physicians integrate information from laboratory reports into their evaluation of a patient’s state.
The steps described in the workflow of radiomics are similar to the steps that need to be considered in the analysis of a blood sample (see Fig.
2).
Just as the drawing and handling of the blood sample may introduce bias,
the acquisition parameters and reconstruction algorithms of a CT scan may as well (11).
Further,
there are specific procedures in the analysis of a blood sample (e.g.,
flow cytometry for the blood count),
and,
in a similar way,
there are different feature selection methods and machine-learning techniques with which to generate a predictive model (12).
The challenge in the implementation of radiomics is the need for rigorous evaluation of every step in the workflow to guarantee the scientific integrity of the extracted imaging feature sets that are to act as a valid biomarker.
What is the benefit of extracting and analyzing large amounts of imaging features rather than a preset few with an underlying pathomechanism already hypothesized?
Predetermining features has the disadvantage of introducing a selection bias by omitting other possibly important features.
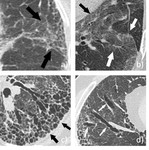
A prominent example can be found in interstitial lung diseases (ILD).
Only the multi-variate information provided by the combination of different CT lung patterns enables the formulation of a working hypothesis.
Even then,
the considerable overlap regarding CT patterns of different ILD entities makes it difficult for radiologists to determine the correct diagnosis (13) (see Fig.
3).
By extracting features from all voxels within the lung,
the probability of gaining additional information can be increased,
which,
when provided to the radiologist,
may increase the confidence in a suspected diagnosis (14).
In theory,
promising sets of features could be hypothesized and evaluated individually.
However,
applying machine-learning methods constitutes a more efficient way,
and allows for the comprehensive extraction and analysis of a “complete” set of imaging features (hence the ending “omics”).
This way,
a predictive set of imaging features (also called a signature (15)) may be found.
Need and Background
As mentioned above,
the primary application for radiomics has been oncology,
with CT being one of the most commonly used imaging modalities,
especially in diagnosing and monitoring diseases of the lung.
Comorbidities in lung cancer,
such as COPD—which affects over 52% of lung cancer patients—have a major impact on survival (4).
By considering these conditions,
and providing a comprehensive evaluation of the changes in imaging features,
predictive risk scores for individual patients can be calculated.
Importantly,
valid and reliable imaging features of such comorbidities retain their importance,
even in the absence of an oncological disease.
They constitute relevant marker patterns across a range of diseases,
and can be studied independently.
A chest CT encompasses the evaluation of all organs of the thorax,
as well as some of the upper abdomen,
and enables the quantitative imaging analysis of important findings (16).
For several common conditions,
possible biomarkers based on quantifiable imaging features were successfully extracted,
warranting an investigation of further in-depth feature extraction:
- COPD—lung parenchyma and airway features (17,
18)
- Osteoporosis—trabecular bone density (19,
20)
- Congestive heart failure—features of interlobular thickening and increased attenuation (21)
- Diffuse interstitial lung diseases—spatial distribution of textural features (22-24)
- Hepatic fibrosis—features of the liver parenchyma (25,
26)
- Cardiovascular disease—features of adipose tissue (27-29)
Relevance of imaging biomarkers in chest CTs using the example of two common diseases:
COPD
In the field of pulmonary diseases,
COPD remains one of the most prevalent and morbid diseases in Europe,
affecting between 4-10% of the adult population and is responsible for approximately €20 billion in direct healthcare costs yearly (30).
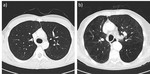
The evaluation of the quantitative detection and evaluation of corresponding imaging features in patients who are referred for chest CTs may open a novel way of risk assessment (see Fig.
4).
As an example,
the PROVIDI study group investigated incidental morphological correlates of COPD on chest CTs acquired for non-pulmonary indications (18).
Emphysema and airway thickening were identified as strong independent predictors of COPD exacerbations that lead to death or hospitalization.
Whereas,
the authors of this work used a visual grading scale for known imaging features,
the automatic quantification of these features is feasible and the subject of ongoing research (31,
32).
Currently,
research in the quantification of the CT imaging correlates of COPD is focused on densitometric estimation of emphysema extent and different methods of airway measurements (such as the full-width-at-half-maximum principle) (33).
However,
there are further abnormalities in lungs that contribute to the state of a patient with COPD,
and which may be quantifiable on chest CTs (e.g.,
mucous plugging and bronchiectasis),
but are not being considered with an approach that focuses on a specific pathological manifestation.
Osteoporosis
Osteoporosis is one of the most common metabolic disorders worldwide and also one of the most common causes for vertebral fractures (34).
Bone mineral density (BMD) is routinely used for the assessment of fracture risk; however,
BMD is only one factor that contributes to vertebral strength (35),
and has limited value in predicting a population at risk (36).
Therefore,
the term “bone quality” was introduced,
which refers to mineralization,
turnover,
and architecture,
and describes not only the absolute amount of BMD,
but also its efficient use (37).
The resulting trabecular bone parameters have been well-established in histological and micro-CT studies.
As an example,
the intra-vertebral heterogeneity of bone density was shown to predict vertebral strength and stiffness,
and consequently,
vertebral failure patterns were explained by local changes in the microstructure (38).
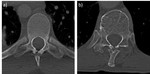
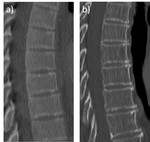
To assess intra-vertebral heterogeneity in a larger population,
features need to be accessible for evaluation in medium- or even low-dose CTs (see Fig.
5 and 6).
Recently,
it was shown that unenhanced chest CTs (19) demonstrated an excellent reliability for the manual quantification of vertebral attenuation values,
and another study demonstrated the potential of low-dose CT for quantitatively evaluating bone fragility and osteoporosis (39).
Challenges
Although much of radiomics research was conducted on data from study patients,
the majority of patients will receive a chest CT in a clinical routine situation (e.g.,
in the field of oncologic imaging,
only 3% of all patients are examined in the setting of clinical trials (40)).
For an accurate,
reproducible,
and valid extraction of imaging features of the lung,
predictors of variation must be considered,
such as scanner-,
protocol- and subject-specific factors (41-43).
Therefore,
one of the main challenges in the field of radiomics is the need for standardized evaluation.
Whereas,
inter-individual variability is not directly controllable and needs to be evaluated for a consequent translation of radiomics to a routine population,
technical parameters are customizable,
with the goal of increasing the reliability of imaging features.
A common conclusion of many studies that have assessed the technical predictors of variability is that these predictors exert a strong influence on imaging features.
However,
certain sets of features often remain stable.
These are several of the known predictors: scanner type (11); dose level (44,
45); slice thickness (8); contrast agent and time after injection (44); and reconstruction algorithms (45,
46).
Inter-scanner reliability:
One study compared the inter-scanner reliability of extracted features between a phantom and lung cancer patients (11).
Whereas,
the variability was the same in vitro and in vivo,
the results demonstrated an overall high variability between different scanners,
with “texture strength” being a relatively stable feature.
Dose level
By comparing different dose levels and reconstruction algorithms,
a diverse susceptibility of features to these conditions was found both in lung nodules and in a uniform water phantom (45).
Interestingly,
smoother reconstruction algorithms could balance out the feature variability at lower dose levels,
further demonstrating the possibility of finding technical settings that would increase the reliability of imaging features.
Contrast agent and time after injection:
In an investigation of the textural features of pulmonary nodules,
a low variation was found in a time window between 60 and 150 seconds after the injection of contrast agent (44).
Reconstruction algorithm:
Shape- and size-based features also showed a good reliability between different reconstruction algorithms (filtered back projection and a Sinogram Affirmed Iterative Reconstruction),
whereas other features (i.e.,
textural features) showed a large variability (46).
While standardizing acquisition parameters and taking advantage of robust feature sets may improve the reliability of results,
it is not possible to account for inter-individual variability in routine data.
Another drawback in data-mining is the statistical demand for large amounts of data.
Many radiomics studies included only small patient datasets,
eventually leading to false-positive results (47).
A way to compensate for these problems is to increase the amount of data.
This,
however,
necessitates the automation of the workflow (e.g.,
segmentation),
which is a cornerstone of the successful implementation of radiomics in routine data,
as will be described in the next section.
Approaches and possible solutions:
Dedicated studies that investigate imaging features regarding their susceptibility to different acquisition parameters will help to find sets of features that remain stable under varying conditions.
For routine imaging data,
this is especially relevant,
as the degree of standardization will be inherently lower than in an experimental setting.
By now,
general measures to increase the quality of radiomics studies can be readily found in the internet,
such as a digital phantom representing a reference feature set (48) and a quality score to assess the scientific integrity of studies (49).
One item of the quality score is robust segmentation,
which should be conducted (semi-) automatically due to the higher accuracy compared to manual segmentation (50).
In addition,
manual segmentation is very time-consuming,
and,
therefore,
cannot be used for large imaging datasets.
After the extraction of features,
selection algorithms identify imaging features with prognostic information regarding the outcome.
The resulting predictive models are based on probabilistic frameworks,
and deep-learning techniques that enable the robust linking of high-dimensional feature spaces to prediction targets.
They are trained on example data and need to be validated on separate datasets,
which again,
requires a large sample size.
Optimizing machine-learning techniques may improve the stability of the prognostic performance of imaging features also during this stage of the workflow.
Chosing the optimal classification method can reduce the performance variation by around 30% (12,
51).
Image repositories:
A comprehensive extraction of hundreds of features, as postulated by the definition of radiomics,
could not only quantify known relationships,
but also find new biomarkers or signatures (52,
53).
For this purpose,
adequate sample sizes are necessary to reduce false discovery rates,
compensate for inter-individual variability,
and provide validation datasets in radiomics studies.
CT-imaging from a clinical population as a data source has the potential of reaching very large sample sizes.
However,
initial research benefits from a higher degree of standardization in study cohorts.
The prognostic value of imaging features found in this way may then be calibrated and validated with large amounts of clinical routine data.
Prognostic models may be transferred from study cohorts to the clinical routine population via transfer learning,
and domain adaptation approaches that exploit a maximum of knowledge from the source domain (e.g.,
cohort) to the target (e.g.
routine),
marking features that are stable,
need adaption,
or are not applicable anymore.
Indeed,
there are a number of publicly accessible CT image databases (54-56),
as well as large study cohorts (57-62) that are highly standardized,
with adequate sample sizes,
and these would be eligible for the high-throughput analysis of quantitative imaging biomarkers (see Fig.
7).
These study cohorts provide available outcome parameters, such as lung cancer diagnosis and stage,
mortality statistics,
cardiovascular events,
and diagnosis of COPD.
Performing assessments on study cohorts or structured data repositories has the advantage of a higher homogeneity of CT data and would greatly add to the practicability of radiomics studies.


![Fig. 2: Images, similar to blood samples, contain data - patients should refrain from eating before blood samples are taken. For some analyses, proper cooling or an additive is necessary (e.g. sodium citrate for blood coagulation parameters). Similarly, patient specific and acquisition parameters may introduce a bias to imaging features in a chest-CT. Following the drawing of blood or the acquisition and reconstruction of an image, a specific method for analysis needs to be chosen. In the case of a blood sample, this may be flow cytometry or haemoglobin electrophoresis. In a CT-image, different sets of features can be extracted from a volume of interest by first applying robust segmentation techniques and then calculating the features with specific mathematical procedures.
At the end of the blood-analysis, the clinician has access to a laboratory report with quantitative information and reference values that aid in clinical decision-making. Radiomics is still a young field of research and the development of quantitative imaging biomarkers for the use in clinical routine poses further challenges. However, in the same manner as the development of laboratory medicine took many years, with the advance of machine-learning techniques and rigorous quality management, a radiology-report including quantitative information from automated feature extraction is achievable. References: Image of hematology analyzer: by HORIBA Medical (Own work) [CC BY-SA 4.0 (http://creativecommons.org/licenses/by-sa/4.0)], via Wikimedia Commons https://commons.wikimedia.org/wiki/File%3APentraDXNexusSPS.jpg](https://epos.myesr.org/posterimage/esr/ecr2017/137601/media/713174?maxheight=150&maxwidth=150)