Random forests for classification were developed keeping the same training and validation sets as for the parametric analysis,
to predict DFS.
Seven different combinations of variables were considered: Clinical (263 patients,
5 features),
CT (295,
41),
PET (258,
43),
PET+CT (258,
84),
CT+Clinical (263,
46),
PET+Clinical (231,
48),
PET+CT+Clinical (231,
89)
Random forests for classification were developed keeping the same training and validation sets as for the parametric analysis.
The outcome to be predicted was the DFS considered until the date of last access or the date of relapse (0=DSF / 1=relapse).
Seven different combinations of variables were considered among clinical predictors and imaging features derived from CT and PET :
• Clinical (263 patients,
5 features)
• CT (295 patients,
41 features)
• PET (258 patients,
43 features)
• PET + CT (258 patients,
84 features)
• CT + Clinical (263 patients,
46 features)
• PET + Clinical (231 patients,
48 features)
• PET + CT + Clinical (231 patients,
89 features)
For each dataset,
a random forest model was built considering different number of trees and different split dimensions.
Moreover,
the relative weight assigned to the output classes was explored.
Once hyper-parameters with a better performance in terms of AUC were identified,
feature importance was extracted for the optimal models.
Additional models were created considering the features with importance greater than the 25th,
50th,
75th and 80th quantiles,
respectively.
The search of the best split dimension was performed again on these new trees.
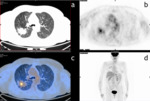
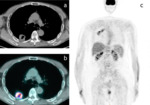
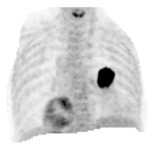
Clinical cases of PET/CT images of a low-risk and a high-risk patient are shown in Fig.4-6 and Fig.5-7.