Keywords:
Pathology, Inflammation, Observer performance, Imaging sequences, Comparative studies, MR-Diffusion/Perfusion, MR, Neuroradiology brain, MR physics, CNS
Authors:
N. Romano1, C. Lapucci2, M. Pardini2, M. Inglese3, L. Roccatagliata2; 1Genova/IT, 2Genoa/IT, 3New York/US
DOI:
10.26044/ecr2019/C-1005
Methods and materials
Patients:
18 patients with MS were enrolled in the study (table 1).

Table 1
MRI imaging analysis:
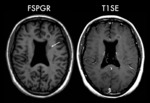
MRI were acquired on a 3T scanner and included T1SE,
FSPGR and TSET2 sequences (3-mm-slice thickness).
Images were analyzed during multiple sessions by consensus of two experienced observers (one neuroradiologist and one neurologist).
Lesion volume (LV) was obtained using manual segmentation technique on TSET2,
T1SE and FSPGR sequences (Jim version 7.0,
Xinapse System).
The lesions number (LN) was counted on T1SE and FSPGR images.
T1SE and FSPGR images were co-registered using a linear registration (NiftyReg package); by subtracting T1SE and FSPGR lesion masks between themselves,
we obtained the lesion masks of the hypointense lesions detectable:
-both on T1SE and FSPGR images (a)
-exclusively on FSPGR images (b)
-exclusively on T1SE images (c)
(a),
(b) and (c) were co-registered using a linear registration to b0 images.

Fig. 2: Example of brain hypointense lesions detected only on FSPGR (red) and on T1SE sequences (yellow), both registered on b0 images.
Diffusion Tensor Images were pre and post-processed to obtain Fractional Anisotropy (FA) and Mean Diffusivity (MD) inside lesions detectable on T1SE and only on FSPGR images were obtained.
Brain parenchymal fraction (BPF) was calculated on FSPGR images.
Statistical analysis:
LV,
LN and FA,
MD were compared between T1SE and FSPGR images using a Student-t test.