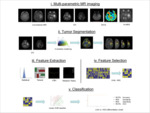
Multiparametric-MRI acquisition and Data post-processing:
Forty patients initially diagnosed with Low- or High-Grade Gliomas (20 LGG & 20 HGG) underwent MRI performed on a 3-Tesla MR whole-body scanner,
applying an advanced imaging examination protocol including,
conventional MRI (T1W-C,
T2W-FSE,
T2W-FLAIR),
MR Spectroscopy (1H-MRS),
Diffusion Tensor Imaging (DTI) and Dynamic Susceptibility Contrast Enhanced MRI (DSCE),
using a 4-channel birdcage and an 8-channel phased-array head coil.
Prior to this retrospective study,
Local Institutional Review Board approval and patient informed consent was obtained (Fig.
1).
FSL software,
was utilized for parametric MR volumes co-registering,
re-slicing into an isotropic voxel size, and applying bias field corrections.
DTI data post-processing was performed with FSL,
including Eddy Current distortions correction,
brain tissue extraction,
Diffusion Tensor estimation and Mean Diffusivity (MD),
Fractional Anisotropy (FA),
Pure Isotropy (p) and Pure Anisotropy (q) parametric maps calculation.
In-vivo SV data analysis and calculation of metabolite ratios were performed with GE Functool software.
Post-processing of the raw spectral data included baseline correction,
frequency inversion and phase shift.
Gaussian curves were fitted to NAA,
Cho,
Cr,
lipid and lactate peaks for determination of peak area.
Finally,
metabolite ratios of NAA/Cr,
Cho/Cr,
ml/Cr and Lipids/Cr were calculated from the area under each metabolite peak for each patient ROI separately.
The Functool software was utilized for processing DSCE data.
CBV maps were calculated for tumors’ representative slices and the mean rCBV measurements were extracted from ROIs placed in areas of maximum contrast enhancement and their contralateral areas.
In addition,
gadolinium uptake time curves were also obtained for identifying the volume of contrast agent maximum uptake for every patient,
which was utilized for textural features extraction.
Volume of Interest (VOI) extraction:
A k-medians clustering segmentation method based on DTI parametric maps, reported in our previous study [10],
was implemented in Matlab 2015b for classifying the brain voxels of each patient into groups with similar isotropic and anisotropic diffusion properties,
accounting for normal and tumorous brain tissue diffusivities.
Specifically,
k-medians clustering (k=16) is applied on a 2D histogram of p (isotropic) and q (anisotropic) components of the diffusion tensor,
derived from all patient cohort.
Subsequently,
RGB color mapping of clusters according to the relative magnitudes of p,
q and T2 values of the cluster centroids and subsequent color assigning to each individual patient’s brain voxels according to their position in the p-q space,
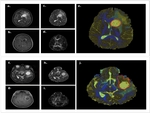
results in whole brain segmented maps (Fig.2).
These color-coded maps are based on the contouring provided by diffusion properties,
being robust in displaying tissue microarchitecture,
thus healthy and tumorous brain tissues present distinctive boundaries.
Finally,
tumor core segments highlighted by the clustering technique,
were delineated on colormaps by an experienced radiologist,
and stacked up to form tumor VOI masks.
Subsequently,
these VOI masks where applied on the various co-registered multiparametric images of our dataset,
resulting in a set of 3D parametric representations of the gliomas.
Radiomic Feature extraction:
Histogram analysis have considered 12 statistical features (minimum,
percentile-25%,
median,
percentile-75%,
maximum,
range,
mean,
standard deviation,
skewness,
kurtosis,
entropy,
uniformity) acquired from normalized data histograms with MATLAB 2015b.
Texture analysis was implemented in MaZda ver.5 software,
considering 11 Gray-Level Co-Occurrence Matrix-based (angular second moment,
contrast,
correlation,
sum of squares,
inverse difference moment,
sum average,
sum variance,
sum entropy,
entropy,
difference variance,
difference entropy) and 5 Gray-Level Run Length Matrix-based features (short run emphasis,
long run emphasis,
gray level non-uniformity,
run length non-uniformity,
fraction),
both calculated on 8-bit quantized images,
and averaged over the 13 3D image directions,
to obtain directionality independence measurements [11].
In summary,
the quantitative radiomic features extracted from the eight 3D tumor parametric VOIs,
regarding p,
q,
MD,
FA,
T1W-C,
T2W-FSE,
T2W-FLAIR and maximum gadolinium uptake volumes of DSCE MRI,
along with the four metabolic ratios of 1H-MRS (Ch/Cr,
NAA/Cr,
mI/Cr,
Lipids/Cr) and mean rCBV values,
resulted in a total of 581 distinct attributes for each subject.
Feature selection and classification:
The machine learning feature selection and classification processes were based on Support Vector Machines (SVM),
implemented in Weka 3.8 software.
Radiomic features were imported in the Support Vector Machine-Recursive Feature Elimination (SVM-RFE) algorithm,
which is a wrapping feature selection method [12].
More specifically,
SVM-RFE iteratively eliminates a set of features by removing the least important one,
according to the weighting vectors of an SVM classifier.
Consequently,
feature ranking in a descending order of discriminative importance is obtained,
according to the elimination sequence.
Afterwards,
the performance of a well-established linear SVM package (SMO) was evaluated with Receiver-Operator-Characteristic (ROC) analysis,
on consecutively SVM-RFE top ranked feature subsets,
to identify the optimal feature subset.
More specifically,
the classifier was repeatedly trained,
starting with the 1 higher-ranked features with a stepwise of adding up 1 feature at each iteration,
and tested with leave-one-out cross-validation. Further investigation through a grid search method aiming in accuracy maximization was utilized for optimizing the hyperparameters of the classification model.