Keywords:
Tissue characterisation, Technology assessment, Segmentation, Computer Applications-Detection, diagnosis, Image manipulation / Reconstruction, CT, Liver, Computer applications
Authors:
A. P. Gonchar, A. Elizarov, N. S. Kulberg, V. Gombolevskiy, I. Blokhin, T. I. Alekseeva, A. Krysanova, M. Suleymanova, D. A. Chernyshev; Moscow/RU
DOI:
10.26044/ecr2019/C-3001
Results
The liver was successfully localized in all cases.
The algorithm requires at least half of true liver volume for accurate segmentation and densitometry.
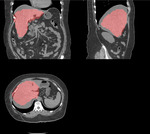
The algorithm correctly identifies the position of the liver even in the following cases:
-
the liver had been scanned partially (which is especially important for chest LDCT)
-
diffuse pathological liver attenuation (i.e.,
in severe steatosis) (Fig.
5)
-
focal liver lesions
The program underestimated real liver dimensions.
We attribute this deviation to shape variations between our model and real samples.
Thus,
improved liver shape database could increase the effectiveness of the algorithm.
Statistical analysis showed Pearson correlation coefficient of 1,
Spearman correlation of 0,903 and rank correlation coefficient of 0,91.
This indicates high accuracy as the segmented area lies well within the borders of real liver.
Difference between manual and automatic measurements was less than 8 HU.