Partecipants
This retrospective analysis was performed in a cohort of men with a clinical indication for prostate biopsy based on prostate-specific antigen elevation and clinical examination or participation in an active surveillance program.
650 patients examined from January 2014 until October 2018 were consecutively imaged with a dedicated prostate mutiparametric MR protocol using an endorectal coil (median age,
67 years; IQR,
50–81 years).
Inclusion criteria were (a) imaging performed on our main institutional 1.5-T MRI system and (b) extended systematic and targeted MRI transrectal US fusion biopsy performed after MRI. Exclusion criteria were (a) history of treatment for prostate cancer (antihormonal therapy,
radiation therapy,
focal therapy,
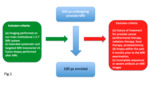
prostatectomy); (b) biopsy within the past 6 months prior to the MRI examination; and (c) incomplete sequences or severe artifacts on MRI images (Fig 1).
On the basis of these criteria,
100 patients were finally included in our study (Figure 1).
MRI protocol acquisition
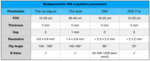
MR images at 1.5 T were acquired prior to biopsy according to the European Society of Urogenital Radiology,
by using the standard multichannel body coil and an endo-rectal coil. Structural and diffusion-weighted MR images (b values = 0,
400,
1200 sec/mm2) were acquired on a 1.5T MR scanner (Table 1).
Imaging findings were scored according the PIRADSv2 by two radiologists in consensus,
blinded to the results of biopsy performed following the imaging session.
MR image processing: lesion segmentation and feature extraction
Three-dimensional volumes of interest (VOIs) of clinical lesions were segmented by one investigator with resident training in prostate mpMRI,
using imaging series and sectional information and pictograms given in the structured clinical report,
in consensus with and under supervision of a board certified radiologist with 5 years of experience in prostate MRI.
Segmentation was performed using a custom Syngo.via Frontiers applicative (SMARTLAB,
IRCCS SDN,
Naples),
and performed on T2- weighted images.
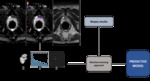
VOIs were drawn on consecutive axial sections by using the polygon tool,
encompassing the whole lesion while avoiding areas of partial volume effects at the border (Fig3).
T2-weighted images,
DWI datasets and ADC maps were coregistered in the same physical space.
Radiomic feature calculations were performed by using a custom matlab code.
Nine shape features (including Number of Voxel,
Maximum and Minimum Diameter,
Volume,
Surface Area,
Surface Volume Ratio,
Compactness,
Spherical Disproportion and Sphericity) were extracted from the tumor segmentation.
First-order and second-order features were extracted from T2 and DWI sequences.
A predictive model was constructed for differentiating biopsy proven prostate cancer (PCa) from non-PCa lesions.
The diagnostic performance of radiomics-based model was compared with that of the PI-RADS v2 scores.
The multivariable predictive model was obtained following the method described by Vallières et al (13),
using at each step an imbalance-adjusted bootstrap resampling (IABR) on 1000 samples.
Finally,
the prediction model was obtained choosing the order that maximize the AUC and computing the final model logistic regression coefficients for the aforementioned combination of feature using IABR.