Purpose
FGT is the volumetric ratio between the fibroglandular tissue and the total breast volume. Its evaluation is a part of the standard breast MR reporting system (BI-RADS). As it is related with mammographic density, it has been recognised as a further risk factor for breast cancer development [1]. Since the MRI is a 3D imaging modality while mammography is 2D, a more accurate evaluation of the volumetric FGT can be performed.
However, breast MR images are characterized by variable contrast depending on the scanner manufacturer,...
Methods and materials
A deep learning approach based on a 3D U-Net architecture was developed to segment breast MR images (Fig. 1). The 3D CNN took one of the multi-modal MRI volumes as input and provided a segmentation mask as output. The whole dataset was split into training (n=88) and test (n=25) set and binary cross-entropy was used as loss function. Dice similarity metric was used to evaluate the quality of the segmentation. The dataset included images of different MRI acquisitions (T1- and T2-weighting, with and without fat-suppression)...
Results
The trend of the loss function and of the Dice index during training are shown in Figure 2. The Dice-Index calculated on validation data set reach maximum value of 0.987 after 380 training epochs. We used this working point to extract FGT maps of our entire population.
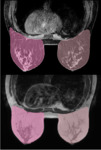
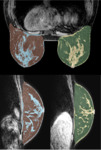
Figure 3 shows an example of the automatic breast segmentation from a multimodal MRI dataset (top image: fat-suppressed Dixon, bottom image: T1-weighted TSE).
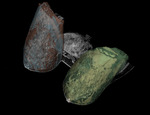
The distribution of the FTG tissue in the left and right breast depicted in axial...
Conclusion
We investigated the feasibility of automatic MRI breast segmentation and FGT map estimation based on a 3D CNN. We achieved good results in a multimodal and multivendor MRI dataset, with Dice Index of 0.987 on unseen volumes.
Limitations: the proposed segmentation method may fail in presence of very large lesions in the breast tissue.
Personal information and conflict of interest
D. della Latta; Massa/IT - nothing to disclose N. Martini; Massa, PLEASE SELECT AN OPTION BELOW/IT - nothing to disclose S. Atzori; Pisa/IT - nothing to disclose D. Chiappino; Massa/IT - nothing to disclose C. Iacconi; Carrara/IT - nothing to disclose V. Piagneri; Carrara (MS)/IT - nothing to disclose T. Trapuzzano; Marina Di Carrara/IT - nothing to disclose
References
[1] King V, Brooks JD, Bernstein JL et al. “Background parenchymal enhancement at breast MR imaging and breast cancer risk.”. Radiology. 2011 Jul;260(1):50-60.