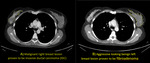
We retrospectively extracted enhanced CT images of 77 lesions (39 malignant versus 38 benign) which are pathologically proven at our institute (King Saud University Medical City) aquired between 2014 to 2018.
All CT images were obtained using (Discovery CT750 HD GE Healthcare, Waukesha, WI) with scanning parameters of 120 kV, 150 mA slice thickness of 3.75 mm( matrix 512 x 512).
Images were reconstructed with filtered back projection technique. All CT images are enhanced post IV contrast obtained at portovenous phase.
All malignant lesions pathologically are invasive ductal carcinoma (IDC) while benign lesions are of variable pathologies as summarized in figure 1.
All lesions were manually traced by third-year radiology resident and validated by 8-year-consultant-radiologist using open source software ImageJ(3)(4) figures 2.
During segmentation, care was taken to include only voxels that resemble vital tumor tissue
We extracted first and second-order statistical features (including 11 Haralick(5)(6) features)(Table 1).
second-order statistical features were averaged for all pixel direction (0, 45, 90 and 135 degrees) figure 3.
WEKA software (7) was used to build a Jrip “Repeated Incremental Pruning to Produce Error Reduction” (8) (RIPPER) classifier with 10-fold cross-validation. This classifier was chosen after feature selection and hyper-parameter optimization automatically generated utilizing Auto-Weka (9).
Auto-WEKA(9), a system designed to help such users by automatically searching through the joint space of WEKA's learning algorithms and their respective hyperparameter settings to maximize performance, using a state-of-the-art Bayesian optimization method using a wide range of feature selection techniques (combining 3 search and 8 evaluator methods) and all classification approaches implemented in WEKA's standard distribution, spanning 2 ensemble methods, 10 meta-methods, 27 base classifiers, and hyperparameter settings for each classifier.
The generated rules of JRIP(RIPPER) classifier are:
-
If (Max <= 121.5) and (Variance <= 3917.3625) and (Contrast <= 341.9005) then CLASS=Benign.
-
if (Min >= 58.422) and (Contrast >= 247.32025) then CLASS=Benign
-
if (Homogeneity >= 0.81025) then CLASS= Benign otherwise
-
CLASS=Malignant