Keywords:
Artificial Intelligence, Lung, CT, Computer Applications-General, Pathology
Authors:
M. M. Pieler, J. Hofmanninger, R. Donner, A. Sikka, E. Jiménez Arroyo, H. Prosch, R. Zhang, G. Langs, A. Makropoulos
DOI:
10.26044/ecr2022/C-15193
Methods and materials
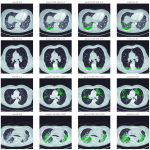
The data annotations for the HC and GGO patterns were carried out by trained annotators on slices from a diverse set of scans that contained the patterns, acquired with different scanners and patient demographics. The annotation process ensured a high quality level through an iterative quality control process. The annotations were reviewed by expert radiologists and re-annotations were carried out until the results were satisfactory. This resulted in 3949 slices that were split into 3074 slices for the training dataset and 875 for the validation dataset.
A subset of the test data with 305 slices was annotated a second time by a different annotator undergoing the same quality control process. This test set was used to estimate the inter-rater agreement for the two different patterns.
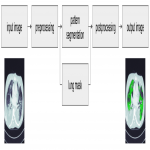
A machine learning segmentation model was trained to segment the HC and GGO patterns. The model architecture was based on a U-Net model which is a widely-adopted deep learning architecture for segmentation tasks in the medical domain [1]. The final model layer outputs a score for the two patterns for every pixel in separate channels. The segmentation model receives a downsampled 256 x 256 2D slice of a preprocessed 3D CT scan as an input and assigns pattern scores to every pixel of the slice. The output segmentation is then post-processed with lung cropping and thresholding of the pattern scores to derive the final segmentation. The general workflow for the pattern segmentation is outlined in Figure 1.
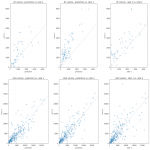
The segmentation model was trained on the annotated lung CT scan slices using data augmentation and a combined loss function. The model training was evaluated based on the dice coefficient [2] on different dataset splits.