Keywords:
Artificial Intelligence, Genital / Reproductive system male, Oncology, MR, MR-Diffusion/Perfusion, Biopsy, Computer Applications-Detection, diagnosis, Cancer
Authors:
N. Debs, A. Routier, B. Lorenzi, L. Wood, F. Nicolas, M.-M. Rohé
DOI:
10.26044/ecr2022/C-22165
Methods and materials
1) Database
- Multi-centric data from various continents (North and South America, Europe, Asia)
- Data acquired with different scanners from different manufactures, with a variety of protocols
- All patients underwent bi-parametric MRI: high b-value Diffusion-Weighted magnetic resonance Imaging (DWI), Apparent Diffusion Coefficient (ADC), T2-weighted (T2w).
- High b-value DWI defined as a DWI with b-value of 2000 if available, otherwise DWI with the closest b-value to 2000 is chosen
- Following bp-MRI, some patients underwent in-bore MRI-guided biopsy or radical prostatectomy. Resulting histopathologic analysis were only available for ~24% of patients.
- All data underwent quality control process
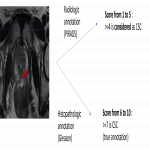
- Based on bp-MRI, PIRADS scores were reported following PIRADS v2.1 guidelines with a single read by a pool of highly experienced radiologists
- Tissue samples from biopsy or prostatectomy were graded with a single read by a pool of using Gleason score (ranging from Benign grade to GS=10). Any suspicious lesion on bp MRI (lesions with PIRADS >=3) was manually delineated on a voxel-level basis.
- Lesions graded PIRADS>3 or GS>6 were considered CSC PCa.
- DWI, ADC, annotated lesion masks registered in T2w space using rigid registration
- Private training/validation data of 2613 patients:
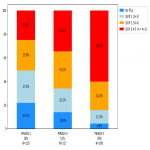
- 2031 patients with PIRADS annotation only (3163 lesions, with 775 lesions with 2<PIRADS<4 and 2388 lesions with PIRADS>=4)
- 562 patients with both PIRADS and Gleason annotations (206 lesions with Benign<GS<7 and 440 lesions with GS>=7)
- Hold-out public data test set: 98 patients with Gleason annotations from ProstateX2 challenge [5] (with 111 lesions: 36 lesions with GS<7, 75 lesions with GS>=7)
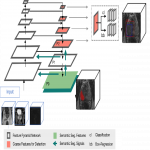
2) Deep Learning model
- Retina U-Net: state-of-the-art method for detection [6]
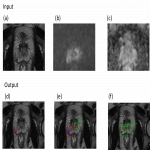
- Localize and classify lesions from given input images (DWI, ADC, T2w). Returning:
- location and classification of lesions by region
- pixel-wise lesion segmentation
3) Experiments
Two distinct experiments were performed:
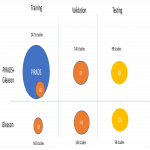
- 1st model trained only on data with GS annotations (training data: 442 patients)
- 2nd model trained on data with PIRADS and GS annotations (training data: 2473 patients)
- PIRADS score = 3 equivalent to GS < 7
- PIRADS score = 4-5 equivalent to GS >= 7
Both models evaluated on same samples for which GS annotation available
- validation data: 140 patients (with 178 lesions: 68 lesions with GS<7, 106 lesions with GS>=7)
- test data: 98 patients from ProstateX2 challenge (with 111 lesions: 36 lesions with GS<7, 75 lesions with GS>=7)
4) Evaluation metrics
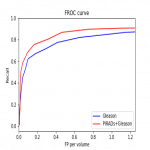
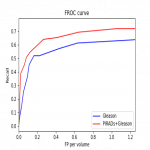
- Detection performances: FROC curve
- Gives the trade-off between the number of detected CSC lesions (GS>=7 lesions) and the number of false positives
- Classification performances: Area Under the Curve (AUC)
- Gives the model ability to classify lesions as either GS>=7 or GS <7