The following steps have been followed to develop the RadSimCT software.
1.1 Development of simulation-based training module on radiation dose optimization for basic training in hardware settings and selecting scan parameters.
To achieve this goal,
we started to develop an interactive and generic CT model for basic training of scan parameters based on real clinical data.
We built,
for DICOM data,
a completely new unproven IT-platform that uses a client-server architecture.
The server stores CT images in a database (up to several TB) using the  Orthanc software.
Metadata containing additional information (radiation doses,
image classification) are stored in
Orthanc software.
Metadata containing additional information (radiation doses,
image classification) are stored in  SQLite databases on the server too.
The client is written in JavaScript and runs in a web browser.
It uses
SQLite databases on the server too.
The client is written in JavaScript and runs in a web browser.
It uses  React and
React and  react-bootstrap JavaScript libraries for the GUI,
react-bootstrap JavaScript libraries for the GUI,
 Redux as the state container and
Redux as the state container and  Cornerstone for the visualization of images (13-18).
CT volumes are transferred to the user’s web browser.
Image processing is done in the web browser via JavaScript.
Cornerstone for the visualization of images (13-18).
CT volumes are transferred to the user’s web browser.
Image processing is done in the web browser via JavaScript.
The server does not process the CT volumes (there is some processing when the CT data are first uploaded to the server).
This allows the new software RSCT to be fully interactive in many common platforms and capable of handling big datasets.
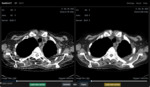
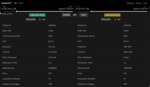
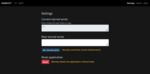
1.2 A generic graphical user interface that can handle data from different vendors has been developed.
A simple user interface that uses generic scan parameters as recommended by the American Association of Physicists in America (AAPM) in joint partnership with several regulatory and industrial entities including the Food and Administration (FDA) (10) is shown in Fig.
1-6.
2. Demonstration of effectiveness of RSCT for enabling radiation dose optimization.
Each output of the project has continuously been validated through end-user participation.
Development of program code has taken place directly in connection with validation sessions with various user groups,
students,
radiologists,
physicists etc. Different usage scenarios for different categories of students,
nurses,
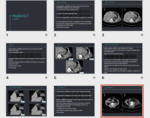
radiologists and physicists have been developed and validated,
see Fig. 7.
Different types of tutorials:
A.
Supervised tutorials:
Students work with RSCT under the supervision of a lecturer.
The lecturer instructs them on what cases to look at (the lecturer provides the Case number) and what further parameters to select (for instance the reconstruction kernel).
The students then experiment with setting the remaining parameters (e.g.
kV and mAs) and observe their effect on the image quality.
B.
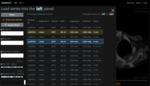
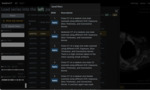
Self-study tutorials:
The user wants to see how an organ (healthy or diseased) looks like on a CT scan.
The user performs a full text search using keywords and the Case description field.
Found cases are listed.
The user selects one and selects other parameters (11).
C.
Reference material
A link (an URL) to a specific patient image in RadSimCT is made from a publication.
When the user follows the link,
RadSimCT displays the specified image (in the left pane) or images (in both panes).
3.
Incorporation of dual-energy CT (DECT) content in the RSCT interface.
A new tool for calculation of Monoenergetic images has been developed (12).
This tool makes it possible to simulate:
- Two-material decomposition to water and iodine
- Monoenergetic image at a photon energy E.
The image is calculated from (i) the known concentrations of water and iodine and (ii) known energy dependence of the mass attenuation coefficient ratio between iodine and water.
This interactive web-based DECT simulation tool will help the user with
- understanding of DECT scanning parameters
- simulation based DECT training exercises
- assessing knowledge of DECT protocols
The main objective of this DECT tool is to understand how to optimize the radiation dose and contrast and increase the clinical value from CT examinations.